巻号一覧

後続誌
37 巻, 5 号
選択された号の論文の12件中1~12を表示しています
- |<
- <
- 1
- >
- >|
巻頭言
-
平野 愛弓2016 年 37 巻 5 号 p. 201
発行日: 2016/05/10
公開日: 2016/05/31
ジャーナル フリーPDF形式でダウンロード (139K)
特集: ナノデザイン表面に基づく生体機能の再構成
-
高久 康春, 鈴木 浩司, 針山 孝彦, 石井 大佑, 森 直樹, 平井 悠司, 下村 政嗣原稿種別: 研究紹介
2016 年 37 巻 5 号 p. 202-206
発行日: 2016/05/10
公開日: 2016/05/31
ジャーナル フリーElectron beam radiation or plasma treatment of the extracellular substances (ECS) of Drosophila's maggots allowed them to survive in the high vacuum environment of a scanning electron microscope (SEM). The plasma-polymerized nano-film of the ECS acts as a protective layer for the rapid dehydration under the reduced pressure of SEM observation. Artificial coating of polysorbitan monolaurate (Tween-20) allowed organisms without a natural ECS to survive under SEM experiment. The results of grazing-incidence small-angle X-ray scattering (GI-SAXS) experiment indicate that the plasma-irradiated surface of the Tween-20 thin film inhibits water evaporation from the insect body. We have proposed a novel method of “NanoSuit” for the high resolution SEM observation of living specimens.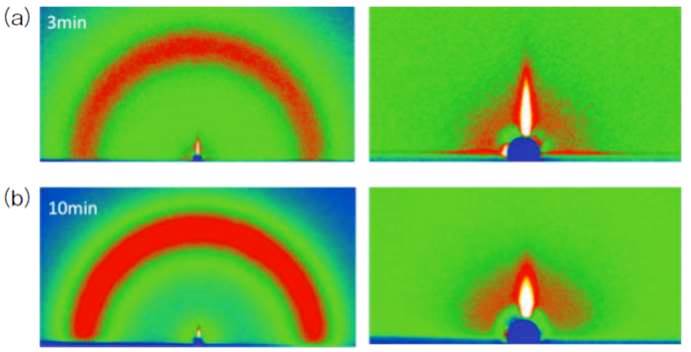 抄録全体を表示PDF形式でダウンロード (561K)
抄録全体を表示PDF形式でダウンロード (561K) -
平口 侑香里, 久代 京一郎, 高井 まどか原稿種別: 研究紹介
2016 年 37 巻 5 号 p. 207-212
発行日: 2016/05/10
公開日: 2016/05/31
ジャーナル フリーIn order to create suitable biocompatible materials for various tissue engineering applications, it is important to be able to understand protein adsorption and cell adhesion behaviors on the material surfaces. It is known that the nanoscale distribution of adsorbed proteins affects cell adhesion behaviors. However, how nanoscale structures affect cell adhesion behaviors is still unclear. Therefore, in this study, we investigated the effect of nanostructures formed by amphiphilic block copolymers on protein adsorption and cell adhesion. Nanoscale phase-separated structures with the different size of the hydrophobic dot-like domains were fabricated by changing the chemical composition, and then the behaviors of protein adsorption and cell adhesion were analyzed. Cells could adhere better on structures with larger hydrophobic dot-like domains because the amount of adsorbed proteins was larger as well. Furthermore, nanoscale phase-separated structures with the opposite polarity were fabricated by changing the solvents. While cells could adhere on structures with the hydrophobic matrix, cells could not adhere on structures with the hydrophilic matrix even though both the amount and density of adsorbed proteins were similar on both structures. This suggests that the simple difference of the distribution of adsorbed proteins can be used to control cell adhesion behaviors. 抄録全体を表示PDF形式でダウンロード (792K)
抄録全体を表示PDF形式でダウンロード (792K) -
安田 賢二, 野村 典正, 寺薗 英之, 金 賢徹, 服部 明弘原稿種別: 研究紹介
2016 年 37 巻 5 号 p. 213-217
発行日: 2016/05/10
公開日: 2016/05/31
ジャーナル フリーTechnologies of analyzing epigenetic information have been developed starting from the twin complementary viewpoints of cell regulation as an ‘algebraic’ system (emphasis on temporal aspects) and as a ‘geometric’ system (emphasis on spatial aspects). Exploiting the combination of latest microfabrication technologies and measurement technologies, which we call ‘on-chip cellomics technology’, we can control and re-construct the interaction of cell-networks from ‘algebraic’ and ‘geometric’ viewpoints. In this review, our developed technologies and some results for spatial viewpoint of epigenetic information as a part of a series of cell-network-based ‘geometric’ studies of celluler systems in our research groups are summerized and reported. The knowlege and technologies acquired from these viewpoints may lead to the use of cells that fully control practical applications like cell network-based drug screening and the regeneration of organs from cells. 抄録全体を表示PDF形式でダウンロード (603K)
抄録全体を表示PDF形式でダウンロード (603K) -
有馬 祐介, 岩田 博夫原稿種別: 研究紹介
2016 年 37 巻 5 号 p. 218-223
発行日: 2016/05/10
公開日: 2016/05/31
ジャーナル フリーEngineering of cell surface with natural and synthetic molecules is expected to provide new approaches for biomedical applications. We have used single-stranded DNA (ssDNA), which was conjugated both to poly (ethylene glycol) and to a terminal phospholipid for cell surface engineering. The lipid moiety is inserted in the lipid bilayer of living cells through hydrophobic interaction, resulting in presentation of ssDNA on the cells surface. This allows for modification of cell surface with biomolecules and attachment of cells to either substrates or different cells, all of which present the complementary ssDNA. The modification of cell surface with proteins or drug-loaded liposomes effectively suppress undesirable biological reactions on cell surface. In addition, high specificity of DNA hybridization and variation of DNA sequences enable to control cell-substrate and cell-cell attachment in a spatially controlled manner. This technique is useful to biomedical applications including cell therapy and tissue engineering. 抄録全体を表示PDF形式でダウンロード (1170K)
抄録全体を表示PDF形式でダウンロード (1170K) -
山本 英明, 平野 愛弓, 谷井 孝至, 庭野 道夫原稿種別: 研究紹介
2016 年 37 巻 5 号 p. 224-229
発行日: 2016/05/10
公開日: 2016/05/31
ジャーナル フリーDissociated neurons form a uniform network in culture that covers the whole coverslip. The number of neurons in the network and extent of their axon/dendrite elaboration can be controlled by using micropatterned surfaces as growth scaffolds. “Defined” neuronal networks thus fabricated make it possible to study how structure of a network correlates with its functional properties. We first describe surface micropatterning techniques can be used to make an array of neurons and to direct axon-dendrite polarity of each cell. To create functional networks, the isolated neurons must subsequently be interconnected. To accomplish this, the cell-repellent domain between individual neurons needs to be altered from cell-repellent to cell-permissive in the culture medium, so that the neurons would be hard-wired. We developed for this purpose a novel surface modification method using titanium dioxide photocatalysis.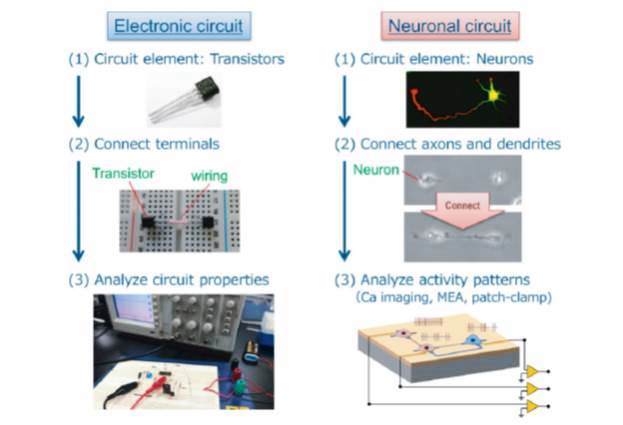 抄録全体を表示PDF形式でダウンロード (1005K)
抄録全体を表示PDF形式でダウンロード (1005K) -
森垣 憲一原稿種別: 研究紹介
2016 年 37 巻 5 号 p. 230-234
発行日: 2016/05/10
公開日: 2016/05/31
ジャーナル フリーBiological membranes play critical roles in the cell. However, the membrane functions remain elusive due to their complex and dynamic nature. We developed a lithographically patterned model biological membrane composed of polymeric and fluid phospholipid bilayers on the solid substrate for mimicking the membrane structures and functions in pre-defined geometries. By modulating the densities of polymeric bilayers, the fluid bilayer spontaneously segregated into liquid-ordered (Lo) and liquid-disordered (Ld) bilayer phases, forming a patterned model system of lipid rafts. Membrane-bound proteins partitioned in the membrane regions according to their affinities to lipid raft. Quantitative evaluation of the interactions between membrane proteins and lipid raft is important to elucidate the functional roles of lipid raft. We also constructed 3D biomimetic structures combining a patterned bilayer and micro-/nano-structures by attaching the polymeric bilayer and an elastomer sheet. Such micro-/nano-spaces would provide a versatile platform for highly selective and sensitive biosensing. 抄録全体を表示PDF形式でダウンロード (1126K)
抄録全体を表示PDF形式でダウンロード (1126K)
連載企画
安全な社会と表面科学
-
廣川 明日菜2016 年 37 巻 5 号 p. 235-237
発行日: 2016/05/10
公開日: 2016/05/31
ジャーナル フリー国立国会図書館は,納本制度に基づき国内出版物の網羅的収集・保存を行う国内唯一の国立図書館である。技術革新に伴い,図書館が取り扱う情報媒体も多様化が進む中,資料の長期保存には様々な課題がある。本稿では当館の取組みを事例に,資料保存の重要性とその課題について整理したい。 抄録全体を表示PDF形式でダウンロード (376K)
抄録全体を表示PDF形式でダウンロード (376K)
談話室
海外研究体験記
-
石井 智2016 年 37 巻 5 号 p. 238-239
発行日: 2016/05/10
公開日: 2016/05/31
ジャーナル フリーPDF形式でダウンロード (322K)
開催報告
-
大門 寛, 長谷川 修司2016 年 37 巻 5 号 p. 240-241
発行日: 2016/05/10
公開日: 2016/05/31
ジャーナル フリーPDF形式でダウンロード (373K)
表面科学技術者資格認定試験例題
-
2016 年 37 巻 5 号 p. 242
発行日: 2016/05/10
公開日: 2016/05/31
ジャーナル フリーPDF形式でダウンロード (1467K)
先端追跡
-
2016 年 37 巻 5 号 p. 243
発行日: 2016/05/10
公開日: 2016/05/31
ジャーナル フリーPDF形式でダウンロード (155K)
- |<
- <
- 1
- >
- >|
