2022 年 19 巻 論文ID: e190015
2022 年 19 巻 論文ID: e190015
DNA mismatches are frequently generated by various intrinsic and extrinsic factors including DNA replication errors, oxygen species, ultraviolet, and ionizing radiation. These mismatches should be corrected by the mismatches repair (MMR) pathway to maintain genome integrity. In the Escherichia coli (E. coli) MMR pathway, MutS searches and recognizes a base-pair mismatch from millions of base-pairs. Once recognized, ADP bound to MutS is exchanged with ATP, which induces a conformational change in MutS. Previous single-molecule fluorescence microscopy studies have suggested that ADP-bound MutS temporarily slides along double-stranded DNA in a rotation-coupled manner to search a base-pair mismatch and so does ATP-bound MutS in a rotation-uncoupled manner. However, the detailed structural dynamics of the sliding remains unclear. In this study, we performed coarse-grained molecular dynamics simulations of the E. coli MutS bound on DNA in three different conformations: ADP-bound (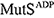 ), ATP-bound open clamp (
), ATP-bound open clamp (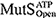 ), and ATP-bound closed clamp (
), and ATP-bound closed clamp (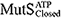 ) conformations. In the simulations, we observed conformation-dependent diffusion of MutS along DNA.
) conformations. In the simulations, we observed conformation-dependent diffusion of MutS along DNA. 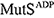 and
and 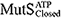 diffused along DNA in a rotation-coupled manner with rare and frequent groove-crossing events, respectively. In the groove-crossing events, MutS overcame an edge of a groove and temporarily diffused in a rotation-uncoupled manner. It was also indicated that mismatch searches by
diffused along DNA in a rotation-coupled manner with rare and frequent groove-crossing events, respectively. In the groove-crossing events, MutS overcame an edge of a groove and temporarily diffused in a rotation-uncoupled manner. It was also indicated that mismatch searches by 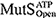 is inefficient in terms of mismatch checking even though it diffuses along DNA and reaches unchecked regions more rapidly than
is inefficient in terms of mismatch checking even though it diffuses along DNA and reaches unchecked regions more rapidly than 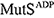 .
.