2015 Volume 90 Issue 4 Pages 231-235
2015 Volume 90 Issue 4 Pages 231-235
CRISPR-Cas9 technology, which uses an RNA-guided nuclease, has been developed as an efficient and versatile genome-editing method to induce mutations in genes of interest. To examine the feasibility of this method in developmental studies of a model monocot, rice (Oryza sativa), we introduced the construct gDL-1, which produced a guide RNA targeting the DROOPING LEAF (DL) gene. DL regulates midrib formation in the leaf and carpel specification in the flower. Because loss of function of DL causes the drooping leaf phenotype in regenerated seedlings, the effect of gene disruption should be easily detected. In transgenic plants carrying gDL-1, the DL gene was disrupted at high efficiency: seven out of nine plants examined had bi-allelic mutations. All transgenic plants with the bi-allelic mutation showed the drooping leaf phenotype. Observation of cross sections of the leaf blade clearly indicated that these transgenic plants failed to make midrib structures, and were comparable to the severe dl mutant dl-sup1. Thus, CRISPR-Cas9 technology can be a useful and efficient tool in developmental studies in rice.
Rapid progress has been made in the understanding of molecular mechanisms that regulate development and morphogenesis in rice (Oryza sativa) during the early 21st century (Hirano et al., 2014; Tanaka et al., 2013, 2014). Important genes have been found in studies using mutants that are defective in various developmental processes. Positional cloning and transposon tagging are efficient methods to isolate genes responsible for morphological defects. For instance, genes critical for shoot apical meristem (SAM) formation during embryogenesis and for floral organ specification in flower development have been isolated by positional cloning (Yamaguchi et al., 2004; Nagasaki et al., 2007). In addition to the forward genetic approach, reverse genetic approaches have become an efficient alternative way to understand the functions of genes of interest. In reverse genetic analysis, mutants that have a defect in the gene of interest are also required. Such mutants have been identified by screening a large number of mutants, in which various genes are disrupted by an insertion of the retrotransposon TOS17 or T-DNA (Hirochika et al., 1996; Jeon et al., 2000). Isolation of knockout mutants by T-DNA is very efficient in Arabidopsis thaliana, because a large number of T-DNA-tagged lines are available worldwide. By contrast, a mutant of interest cannot be easily obtained in rice, because of the limited number of tagged lines. Recently, the TILLING method (Suzuki et al., 2008; Satoh et al., 2010) has become available in rice. Although this method is helpful to obtain mutants, the small size of mutant pools is also a limiting factor to isolate mutants of interest in rice.
The recently developed RNA-guided endonuclease CRISPR-Cas9 (clustered regulatory interspaced short palindromic repeats/CRISPR-associated protein 9) system is an efficient and versatile technology to modify endogenous genes of interest in various organisms (Jinek et al., 2012; Cong et al., 2013; Li et al., 2013; Mali et al., 2013; Nekrasov et al., 2013; Shan et al., 2013; Wang et al., 2013). This technology is based on a bacterial immune system to inactivate foreign DNAs such as virus DNA (Harrison et al., 2014; Hsu et al., 2014; Terns and Terns, 2014). The CRISPR-Cas9 system has several advantages over other genome-editing technologies using artificial nucleases such as TALENs (transcription activator-like effector nucleases) and ZFNs (zinc-finger nucleases). First, it is easier to make a construct to disrupt the gene of interest using CRISPR-Cas9 technology. Second, the efficiency of gene disruption is very high. Third, it is easy to obtain bi-allelic mutations in regenerated plants without having to make the M2 generation.
To test the efficient production of developmental mutants using CRISPR-Cas9 in rice, we focused on the DROOPING LEAF (DL) gene, which is required for midrib formation in the leaf and carpel specification in the flower (Yamaguchi et al., 2004). The midrib is a strong structure formed in the central region of the leaf and is responsible for erectness of the leaf blade in rice (Ohmori et al., 2008, 2011). DL positively regulates midrib formation by promoting cell proliferation along the adaxial-abaxial axis in the central region of the primordia. The dl mutant fails to proliferate cells along this axis and to form the midrib, resulting in a drooping leaf phenotype. Because this leaf phenotype is easy to distinguish from the wild-type leaf phenotype, we chose the DL gene to disrupt using CRISPR-Cas9 technology as a model experiment in a developmental study. In this paper, we describe the efficient production of DL knockout lines and the resulting morphological phenotypes of those mutants.
To obtain knockout lines of the DL gene, we made a construct using a gRNA/Cas9 all-in-one vector, pZH_OsU3gYSA_MMCas9, in which the Cas9 gene and gRNA are driven by the double cauliflower mosaic virus 35S promoter and the OsU6 promoter derived from the rice U6-2 snRNA gene, respectively (Mikami et al., under submission; Feng et al., 2013). The 20-bp sequence shared by the first intron and second exon of DL was selected as the target of guide RNA (Fig. 1A). After annealing of synthetic oligonucleotides including the target sequence and 5’ ends adaptor sequence, the resulting double-stranded DNA fragment was first inserted into the BbsI restriction site of a gRNA cloning vector, pU6gRNA-oligo, which has the OsU6 promoter and gRNA scaffold (Mikami et al., under submission). The OsU6 promoter and gRNA region was then excised from the gRNA cloning vector, and introduced into the AscI and PacI sites of pZH_OsU3gYSA_MMCas9. The resulting construct, named gDL-1, was introduced into rice calli by Agrobacterium-mediated transformation (Hiei et al., 1994). Calli in group A were cultured in selection medium for three weeks, whereas those in group B were cultured for five weeks. After this hygromycin selection, calli were transferred to regeneration medium, and transgenic rice plants were regenerated (plants A1–A5 from group A calli; plants B1–B4 from group B calli).
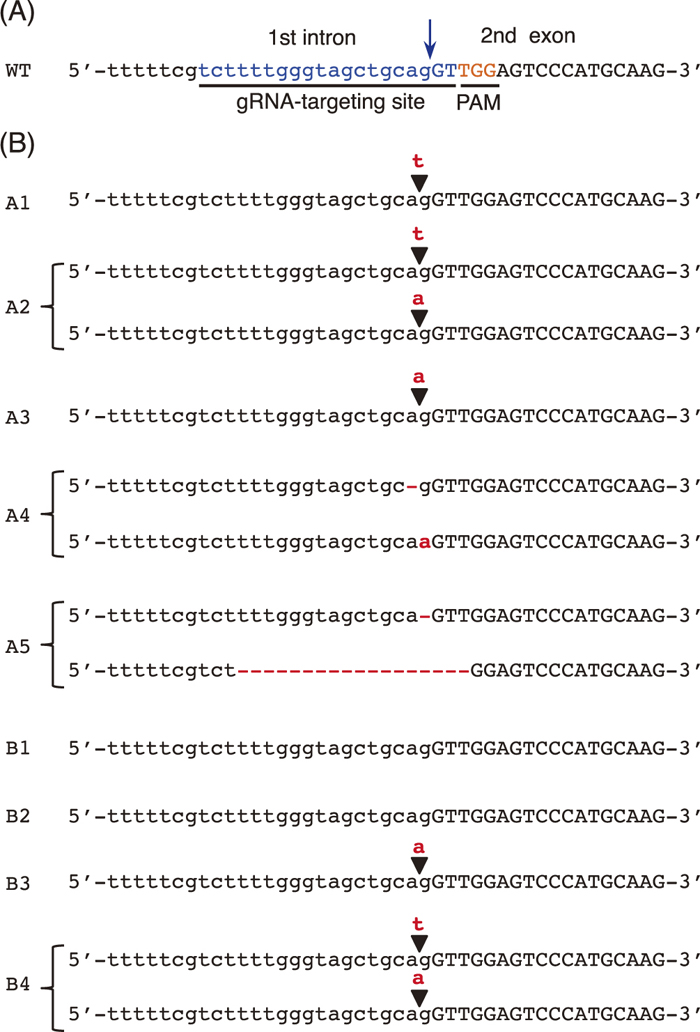
Nucleotide sequences including the target site of the guide RNA in the DL gene. (A) Nucleotide sequence of DL in wild type. (B) Mutated and non-mutated nucleotide sequences in each transgenic plant carrying gDL-1. One sequence in a transgenic line means that the same mutation occurs in both alleles. The nucleotide sequences in the first intron are indicated in small letters, whereas those in the second exon are indicated in capital letters. The junction of the first intron and the second exon is indicated by an arrow. Arrowheads and dashes indicate insertions and deletions, respectively. Mutated nucleotides and deletions are shown in red. A1, A2, A3, A4 and A5 plants were regenerated after 3 weeks of growth on the selection medium, whereas B1, B2, B3 and B4 plants were regenerated after 5 weeks of growth.
Genomic DNA was isolated from each transgenic plant, and the nucleotide sequence of the target site in the DL gene was determined. Except for two plants (B1 and B2), seven of the nine regenerated plants (A1, A2, A3, A4, A5, B3 and B4) had a mutation at the target site (Fig. 1B). In many cases (10 out of 14 alleles), a single nucleotide was inserted at the same position near the junction between the first intron and the second exon. In two other cases, a single nucleotide was deleted from similar positions. A single nucleotide substitution and an 18-nucleotide deletion were also observed. When mutation occurred, all plants had bi-allelic mutations. Thus, CRISPR-Cas9 methods were highly efficient in disrupting the rice DL gene.
Rice leaves have a strong structure, called the midrib, in the central region, and stand upright due to this structure (Fig. 2A). Because DL plays a crucial role in forming the midrib (Yamaguchi et al., 2004; Ohmori et al., 2011), the loss-of-function mutant of DL (dl-sup1) lacks the midrib, resulting in a drooping leaf phenotype (Fig. 2B).
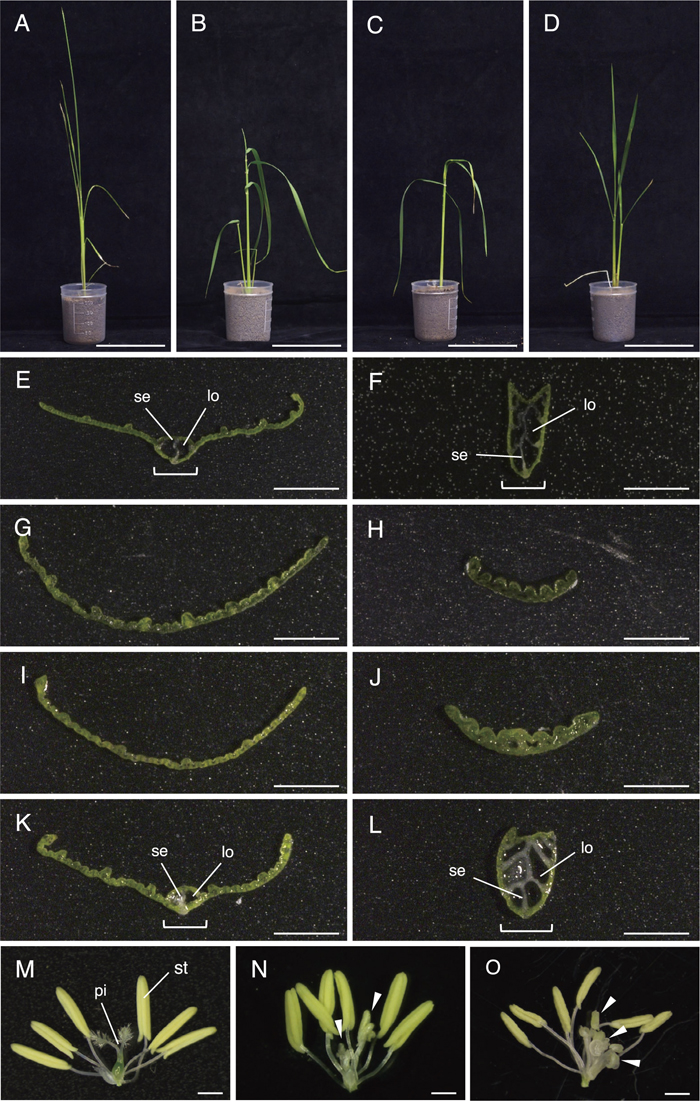
Phenotypes of leaves and flowers. (A–D) Seedlings of wild-type (A), dl-sup1 (B), and transgenic plants A1 (C) and B1 (D). (E–L) Cross sections of the leaf blade in wild-type leaf (E, F), dl-sup1 (G, H), and transgenic plants A1 (I, J) and B1 (K, L). (E, G, I and K) the middle part of the leaf blade; (F, H, J and L) the proximal part of the leaf blade. (M–O) Flowers of wild type (M), dl-sup1 (N) and the transgenic plant B3 (O). Brackets and arrowheads indicate the midrib and ectopic stamens, respectively. lo, locule; pi, pistil; se, septum; st, stamen. Bars = 10 cm in (A–D); 1 mm in (E–O).
We first observed the phenotypes of the seedlings after regeneration. Transgenic plant A1, which had a bi-allelic mutation in DL, showed the drooping leaf phenotype, similar to that observed in the dl-sup1 mutant (Fig. 2, B and C). Similarly, all other plants with a bi-allelic mutation showed the drooping leaf phenotype (Table 1). By contrast, transgenic plants B1 and B2, which had no mutation, showed an upright leaf phenotype, like wild type (Fig. 2, A and D; Table 1). Thus, genotypes were consistent with phenotypes in all transgenic plants examined.
| plant | A1 | A2 | A3 | A4 | A5 | B1 | B2 | B3 | B4 |
|---|---|---|---|---|---|---|---|---|---|
| bi-allelic mutation | yes | yes | yes | yes | yes | no | no | yes | yes |
| phenotype | dl | dl | dl | dl | dl | WT | WT | dl | dl |
We next examined cross sections of the leaf blade. In the middle part of the wild-type leaf, a midrib structure consisting of two locules and a septum was observed (Fig. 2E), as shown in previous papers (Yamaguchi et al., 2004; Ohmori et al., 2011). In the proximal part (Fig. 2F), the midrib was enlarged, and was composed of several locules and septa. By contrast, no such structure was observed in either the middle or the proximal part of the leaf blade in dl-sup1 (Fig. 2, G and H). Transgenic plant A1 lacked the midrib in both the middle and proximal part, consistent with the bi-allelic mutation in DL (Fig. 2, I and J). Transgenic plant B1with no mutation showed normal formation of the midrib (Fig. 2, K and L).
Finally, we examined flower phenotypes, because DL is required for carpel specification (Yamaguchi et al., 2004). Rice flowers have two lodicules, six stamens and one pistil, which is formed from three carpels (Fig. 2M; Yamaguchi et al., 2004). Carpels are homeotically transformed into stamens in the dl-sup1 mutant (Fig. 2N). To examine flower phenotypes, we used the transgenic plant B3, which also showed the drooping leaf phenotype. In this transgenic plant, carpels were not formed, and many ectopic stamens were formed instead of them (Fig. 2O). We observed 18 flowers in this line. All flowers examined exhibited homeotic transformation of the carpel into the stamen. Thus, carpel specification was also disturbed by the disruption of DL by the CRISPR-Cas9 system.
In this paper, we generated several mutants in which the DL gene was disrupted by CRIPR-Cas9 technology, and showed that the mutants exhibited the morphological phenotypes expected from the loss of function of the DL gene. The efficiency of introducing mutations was very high using this technology: seven out of nine regenerated plants had bi-allelic mutations. The efficient production of bi-allelic mutations should provide useful materials to understand developmental genes of interest, because phenotypes can be analyzed using regenerated plants per se without the need to produce homozygous mutants by self-pollination. Rice has a relatively long life cycle, as compared with Arabidopsis. Therefore, the production of null mutants in a short period is a great advantage in rice research. In addition, multiple genome engineering has already been achieved by this CRIPR-Cas9 technology (Cong et al., 2013; Li et al., 2013; Mali et al., 2013; Wang et al., 2013). It should also be feasible to make a multiple mutant within a few months in rice.
All mutations occurred near the splicing acceptor site of the first intron. According to the GT-AG rule of splicing in protein-coding genes (Breathnach and Chambon, 1981; Mount, 1982), the disruption of the acceptor sequence “AG” seems to cause serious defects in splicing of the DL transcripts and in producing normal DL protein, probably because of premature termination in the first intron. If an “A” nucleotide is inserted in this acceptor sequence (Fig. 1, A and B (A2 lower allele, A3, B3, and B4 lower allele), the sequence would still be “AG” at this site. This suggests that the acceptor sequence remains intact. However, all four of these transgenic plants (A2, A3, B3 and B4) showed a severe drooping phenotype, like the dl-sup1 mutant. Therefore, a sequence other than “AG” at the splicing junction may be necessary for proper splicing of this intron.
This research was supported in part by Grants-in-Aid for Scientific Research from MEXT (23248001, 25113008 to H.-Y. H.).