2017 Volume 92 Issue 5 Pages 223-234
2017 Volume 92 Issue 5 Pages 223-234
Proteins belonging to the sigma factor family in eubacteria initiate transcription by associating with RNA polymerase. A subfamily, the extracytoplasmic function (ECF) sigma factors, which form a widely distributed bacterial signal transduction system comprising a sigma factor and a cognate membrane-embedded anti-sigma factor, regulates genes in response to stressors that threaten cell envelope integrity including the cell wall and membrane. The Gram-positive soil bacterium Bacillus subtilis provides a valuable model for investigation of the ECF sigma factors. This review focuses on the function and regulation of ECF sigma factors in B. subtilis, in which anti-sigma factors play a role in connecting an external stimulus with gene regulation. As representative examples, the regulon and regulatory mechanism of σW are closely associated with membrane-active stressors, whereas σM is strongly induced by conditions that impair peptidoglycan synthesis. These studies demonstrate that the mechanisms of ECF-dependent signaling are divergent and constitute a multi-layered hierarchy, and provide useful insights into the elucidation of unknown mechanisms related to ECF sigma factors.
Bacteria are roughly classified into two groups, enteric bacteria that reside within the animal or human gut and environmental bacteria that live in natural habitats globally. The former situation is relatively stable because the bacteria depend on their host environment, which is under homeostatic control. In contrast, the latter situation ranges from benign to extremely hostile surroundings and can change dynamically and suddenly. In order to adapt to and survive such hostile conditions, bacterial cells have developed complex stress response mechanisms. The surface of a cell is directly exposed to the external environment, and functions as a rampart to protect the cell interior. The cell surface also acts as a sensor, which perceives the external state and transduces the information across the surface, thus allowing adaptation. Through this process, physical information (such as external stressors) is converted to biological signals, which causes alteration of gene expression, fundamentally at the transcriptional level. The cell surface of free-living bacteria is basically composed of a cell wall and membrane. The sensing machinery, in many cases, resides within the cell membrane as membrane-embedded proteins.
The genes required for stress responses are not constantly transcribed during cell growth. In bacteria, transcription initiation is regulated almost entirely by sigma factors, which are components of the RNA polymerase complex. Sigma factors recognize specific promoter sequences and allow genes to be transcribed. A particular group of sigma factors is activated only when they are needed for a particular response, resulting in directed gene transcription specific to the response. In this regard, some sigma factors are regulated by cognate anti-sigma factors, which impedes RNA binding, and the so-called ECF (extracytoplasmic function) sigma factors are representative of such sigma factors. The ECF subfamily was first designated in a study on Streptomyces coelicolor σE, a sigma factor controlling the extracellular agarase gene (Lonetto et al., 1994). Characteristically, σE of S. coelicolor lacks several domains that are conserved among the major family of sigma factors, and its structurally homologous sigma factors also regulate functions related to the cell surface response to extracytoplasmic stimuli. To date, advances in genome sequencing have revealed that environmental bacteria especially possess multiple ECF sigma factors and that numerous sigma factors expressed by diverse bacteria are categorized in the ECF subfamily. The anti-sigma factor is a membrane-embedded protein composed of an external domain displayed on the cell surface and an internal domain located within the cell. The internal domain binds a cognate sigma factor and thus sequesters it from RNA polymerase. The external domain senses an environmental signal and transmits it to the internal domain, resulting in the release of the sigma factor, which can then bind to RNA polymerase. Although many ECF sigma factor-anti-sigma factor pairs are found in genome databases, little is known about the molecular mechanism of regulation of ECF sigma factors by their anti-sigma factors - for instance, how anti-sigma factors receive and sense signals, and how they release the sigma factors.
One of the commonest Gram-positive and environmental soil bacteria, Bacillus subtilis, has been studied for a century as a model organism in many fields of biology. When the surrounding nutrition is exhausted, these bacteria produce a spore within the cell, which is highly dehydrated, tolerant to long storage and resistant to unfavorable conditions. Additionally, B. subtilis possesses response mechanisms for adapting to various environmental stresses such as high temperature, high osmolality and oxidative conditions. Bacillus subtilis has seven genes that encode ECF sigma factors, of which six are apparently regulated by individual anti-sigma factors as the genes encoding ECF sigma and anti-sigma factors constitute an operon (Fig. 1); these six are also involved in similar mechanisms for stress response (Table 1). An important approach to unraveling the cellular function of sigma factors is to identify the genes whose transcription is under direct control of these sigma factors, using comprehensive analyses such as microarray analysis (Asai et al., 2003), and many other investigations have been undertaken to date. A highly detailed and general description about the structure and function of ECF sigma factors is provided in three recent reviews (Mascher, 2013; Paget, 2015; Helmann, 2016). In this issue, I focus on regulatory aspects of anti-sigma factor activity, especially based on the cell surface (i.e., cell membrane and cell wall) dynamics, mainly in B. subtilis, compared to the representative model proposed in other bacteria.
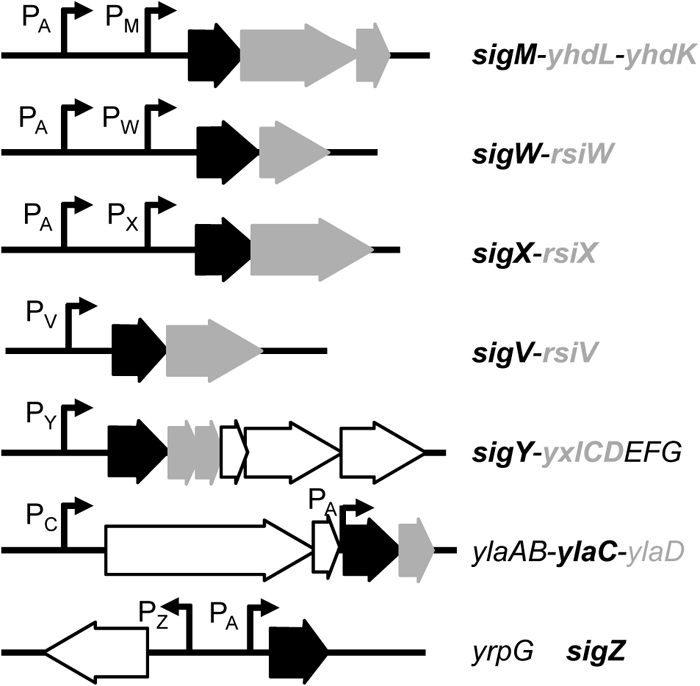
Gene organization around the ECF sigma factor genes. The ECF sigma factor genes and their cognate anti-ECF sigma factor genes are indicated in black (arrows and bold letters) and gray (arrows and bold letters), respectively. The locations of promoters (P) are indicated with hooked arrows, and the subscript after ‘P’ indicates the last letter of the sigma factor that recognizes the promoter. The subscript ‘A’ indicates SigA, a housekeeping and essential sigma factor in B. subtilis.
| σ | Anti-σ | Activation stress signals | Function | Reference |
|---|---|---|---|---|
| σ A | – | – | housekeeping | |
| σ M | YhdL/YhdK | salt, cell wall antibiotics, ethanol, heat, acid, superoxide stress | cell wall homeostasis | Thackray and Moir (2003) |
| σ W | RsiW | membrane-active agents, alkali | cell membrane homeostasis | Helmann (2002) |
| σ X | RsiX | cationic antimicrobial peptide | cell surface homeostasis | Helmann (2002) |
| σ V | RsiV | lysozyme | lysozyme resistance | Ho et al. (2011) |
| σ Y | YxlC/YxlD | nitrogen starvation | unknown | Tojo et al. (2003) |
| σ YlaC | YlaD | oxidative stress | unknown | Matsumoto et al. (2005) |
| σ Z | unknown | salt | unknown | Asai et al. (2005) |
| σ B | RsbW | energy stress, environmental stress | general stress response | Hecker et al. (2007) |
Nineteen sigma factors are encoded in the B. subtilis genome; among these, seven belong to the ECF sigma factor subfamily and are called σM, σV, σW, σX, σY, σZ and σYlaC (Table 1). Figure 1 shows their gene organizations with the location of their promoters and their anti-sigma factor genes. The genes for a sigma factor and its anti-sigma factor constitute an operon in most cases. Generally, the N-terminal portion of the anti-sigma factor, which is flanked by the predicted membrane-spanning region, resides in the cytosolic space and interacts with the partner sigma factor. In the case of RpoE and RseA, which are an ECF sigma factor and its anti-sigma factor, respectively, in Escherichia coli, the cytosolic domain of RseA was revealed to be sandwiched between the so-called region 2 and region 4 domains of RpoE, based on crystal structure analysis, thus blocking the interaction between RNA polymerase and the sigma factor (Campbell et al., 2003). In contrast, with the cobalt and nickel resistance-regulating ECF sigma factor CnrH and its anti-sigma factor CnrY in the β-proteobacterium Cupriavidus metallidurans, the cytoplasmic domain of CnrY embraces the region 2 and region 4 domains of CnrY, thus blocking RNA polymerase from promoter recognition (Maillard et al., 2014). The direct interaction of sigma factors with their anti-sigma factors in B. subtilis was examined using a yeast two-hybrid system (Yoshimura et al., 2004). This analysis showed that the sigma factors bind their cognate anti-sigma factors exclusively, indicating that the interaction of each pair is highly specific. The amino acid sequences of anti-sigma factors are evolutionarily unrelated (although well-conserved counterparts are found among closely related species) (Helmann, 2002). This evoked the idea that the pairs of a sigma factor and an anti-sigma factor might have co-evolved and been inherited through vertical transfer. This notion also correlates with the fact that the environmental signals to which the anti-sigma factors respond are not unified even though they sometimes overlap, and the regulatory mechanisms that control their function, most of which are not uncovered, are potentially diverse.
Typically, the transcription of each operon is regulated by the sigma factor encoded within the operon (Fig. 1). Therefore, the activities of sigma factors are measured by the strength of transcription directed by promoters that they themselves recognize. Basically, disruption of the anti-sigma factor leads to activation of the cognate sigma factor (Asai et al., 2003). The effect of disruption of the anti-sigma factor genes except for yhdL-yhdK and yxlC-yxlD on several promoters was tested in a round-robin system (Table 2). Inactivation of yhdL-yhdK and yxlC-yxlD, the anti-sigma factors of σM and σY, respectively, was not successful, suggesting that abnormally high expression of σM and σY is toxic to the cell (Horsburgh and Moir, 1999; Tojo et al., 2003). Although the promoter sequences recognized by ECF sigma factors are very similar, as shown in Fig. 2, promoter activity increases only when the corresponding sigma factor is activated by deletion of its cognate anti-sigma factor gene. Simultaneous induction of multiple ECF sigma factors by a single ECF sigma factor induced with a stimulus to the cell seems to be avoided. As mentioned above, these observations support the idea that each sigma factor might have emerged independently even though ECF sigma factors are categorized as one large subfamily of sigma factors based on their structural similarities.
| Mutation | Promoter fused with lacZ gene | |||||
|---|---|---|---|---|---|---|
| PsigM | PsigV | PsigW | PsigX | PsigY | PylaA | |
| Wild | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
| rsiV− | 0.79 | 248 | 0.41 | 0.63 | 0.94 | 4.14 |
| rsiW− | 0.72 | 0.87 | 2.08 | 0.85 | 0.53 | 1.54 |
| rsiX− | 0.70 | 0.84 | 0.59 | 1.26 | 0.62 | 2.20 |
| ylaD− | 0.79 | 0.53 | 0.77 | 0.96 | 0.52 | 1.42 |
The activities of β-galactosidase are shown as relative values compared to the activity in wild type. Figures in bold indicate activities caused through self-regulation, as determined in previous studies.

Promoter sequences of the ECF sigma factor genes. The first five promoters are located just upstream of the sigma factor genes and are recognized by the sigma factors themselves. In the last two promoters, ylaA is the first gene of the operon including ylaC encoding for ECF sigma factor σYlaC. This promoter is recognized with σYlaC. The gene yrpG is located upstream of sigZ (encoding the ECF sigma factor σZ), in the opposite orientation, and its promoter is recognized by σZ.
The signals to which ECF sigma factors respond tend to reflect their function. For instance, transcription of sigV is induced by lysozyme, and σV induces genes that confer lysozyme resistance to the cells (Ho et al., 2011). The operon containing the sigY gene is induced under nitrogen-starved conditions, but the physiological significance of this sigma factor is not clear (Tojo et al., 2003). σYlaC resides within an operon comprising four genes, in which transcription of ylaC-ylaD is induced when cells are exposed to oxidative stress dependent on Spx (Matsumoto et al., 2005). No detailed study on σZ, which is an orphan sigma factor (the anti-sigma factor is unknown), has been performed to date. In contrast, σM, σW and σX are expressed to some extent during normal growth and, together with σV, their function and regulatory mechanism have been well studied. These three or four sigma factors can play crucial roles in stress responses in B. subtilis (Asai et al., 2005; Helmann, 2016), acting like “The Three Musketeers or Four”. Each of the seven ECF sigma factors is individually dispensable for growth (Kobayashi et al., 2003), but they are involved in response mechanisms to various external stresses. Moreover, these seven genes can also be simultaneously deleted from the genome (Asai et al., 2008; Luo et al., 2010). Although the regulatory mechanisms comprising the sigma factor and anti-sigma factor are diversified, the signals that stimulate these ECF sigma factors in B. subtilis and their regulon genes partly overlap (Mascher et al., 2007), indicating that they fight cooperatively against external stress. The northern blots in Fig. 3A show the transcripts of indicated genes after cells were exposed to a high temperature for 30 min. The ypuA gene was induced by heat treatment (lanes 1 and 6). This induction was weakened by inactivation of sigM (lane 7) and almost abolished by simultaneous deletion of sigM and either sigW or sigX (lanes 9 and 10), suggesting that these three sigma factors are involved in the transcription of ypuA. In contrast, double deletion of sigM and sigV or sigX (lanes 8 and 10) restored the downshift of the expression of yjbC and ywoA to the level of the sigM+ strain, and this restoration was abolished in the sigW− strain (lane 9). These results suggest that σW, which is not drastically induced by heat in wild type cells, was activated to cope with the crisis in the absence of σM, σX and σV. Thus, the role of σW in heat stress may be obscured by σM, σX and σV. Notably the expression of sigW was constitutively higher than that in the wild type strain when the six ECF sigma factor genes other than sigW were deleted and σW was the sole ECF sigma factor in B. subtilis (Fig. 3B). This suggests that the cells perceive a crisis from the state of less responsive mechanisms and exert the best possible measures to overcome it using their remaining resources. The robustness of cells against stresses that threaten their existence may have been accomplished through such multilayered and overlapping mechanisms (Radeck et al., 2017).
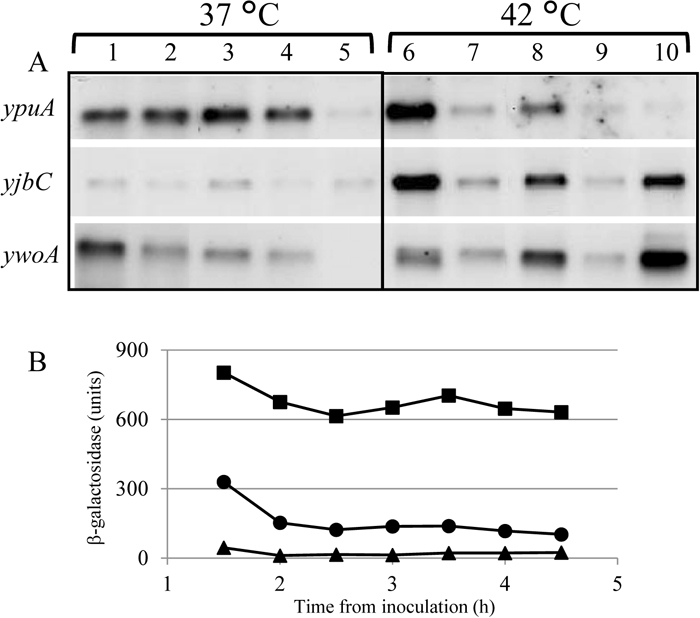
Effect of sigma factor deletion on the activity of the remaining sigma factors. (A) Northern blotting analysis visualized using probes hybridizing with the genes indicated on the left. Total RNA was extracted from the cells before heat treatment (37 ℃) (lanes 1 to 5) and after heat treatment (42 ℃) for 30 min (lanes 6 to 10). The genotypes of the strains used in this experiment are as follows. Lanes 1 and 6: parental strain (B. subtilis 168); lanes 2 and 7: sigM−; lanes 3 and 8: sigM− and sigV−; lanes 4 and 9: sigM− and sigW−; and lanes 5 and 10: sigM− and sigX−. (B) The cells harboring the promoter recognized by σW fused to lacZ (Kosono et al., 2004) were harvested at the indicated times and β-galactosidase activity was measured. Circles: wild type; squares: sigM−, sigV−, sigX−, sigY−, sigZ− and ylaC−; triangles: sigM−, sigV−, sigW−, sigX−, sigY−, sigZ− and ylaC− (Asai et al., 2008).
The environmental signals that activate ECF sigma factors through the anti-sigma factors are expected to stimulate the cells physically and modulate the proteins related to the ECF sigma factor-responding system. The best-studied mechanisms that regulate the intracellular behavior of anti-sigma factors are observed with σW and RsiW (Fig. 4). After exposure of cells to stress conditions, the transmembrane segment of RsiW is cleaved at site 1 near the boundary of the extracellular region by a membrane-embedded protease, PrsW (protease responsible for activating sigma-W; formerly YpdC) (Ellermeier and Losick, 2006), and is then cleaved at site 2 near the boundary of the intracellular region by another membrane protease, RasP (regulating anti-sigma factor protease; formerly YluC) (Schobel et al., 2004). The intracellular portion (anti-sigma factor binding) released through this proteolytic action is eventually degraded by the ClpXP protease complex (Zellmeier et al., 2006), resulting in the complete disengagement of σW from RsiW. This mechanism is similar to the regulation of RseA, the anti-sigma factor of σE in E. coli, by DegS and RseP proteases (Ades et al., 1999; Alba et al., 2002; Kanehara et al., 2002). This sequential cleavage of a protein within a membrane is called regulated intramembrane proteolysis (RIP) and was first described in the regulation of mammalian cholesterol metabolism by proteolysis of the membrane-bound transcription factor SREBP, which is also cleaved sequentially by two membrane proteases (Brown and Goldstein, 1997). In contrast to the fact that the cleaved portion of the anti-sigma factor is not useful for cellular activity, SREBP is a precursor that is cleaved to its active form as a transcriptional factor. The principle of RIP is employed in two different types of such transcriptional regulation in mammals and bacteria. This fact evokes the idea that the bacterial two-component-type system, composed separately of a transcriptional factor and a regulatory anti-factor, developed into a one-component-type system through fusion. ECF sigma factors classified as group ECF41 contain a C-terminal extension, which seems to be a fused anti-sigma-like domain and is also required for sigma factor activity (Wecke et al., 2012). The activity of CorE-like sigma factors classified as group ECF44 is also positively or negatively regulated by a C-terminal extension (Gómez-Santos et al., 2011). A putative ECF sigma factor gene prolonged with a putative anti-sigma factor gene in a single open reading frame is occasionally found in the bacterial genome database (e.g., OB_RS12020 in Oceanobacillus iheyensis HTE831).
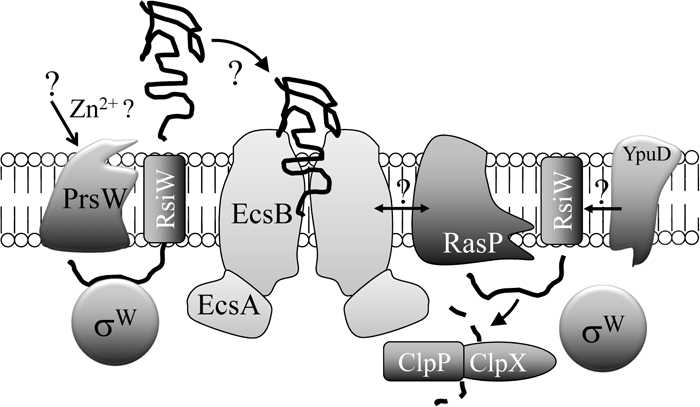
Model of the mechanisms of RsiW degradation resulting in σW activation in B. subtilis. The details are described in the text. Still-unresolved function, interaction, signal and mechanism are indicated by question marks.
When RasP was the only known protease that regulated the proteolysis of RsiW in B. subtilis, many studies were undertaken to identify a protease functioning upstream of RasP. We tested the effect of inactivating all the annotated protease genes encoded in the B. subtilis genome on σW-dependent transcription. As PrsW was a novel family of membrane proteases and was not annotated as a protease at the time, we overlooked it. However, we found that inactivation of ypuD, encoding a protein of unknown function, led to a slight decrease in σW activity (K. A., unpublished results). Sequence similarity searches indicate that YpuD is a putative intramembrane metalloprotease with four transmembrane domains and is a member of the so-called CPBP (CAAX proteases and bacteriocin-processing enzymes) family, a large group of diverse proteins including the type II CAAX prenyl endopeptidases (Pei and Grishin, 2001). The involvement of YpuD in the regulation of RsiW has not yet been analyzed in detail.
Other investigations using transposon-mediated random mutagenesis were performed to elucidate the unknown regulatory factors of RsiW, including still-undiscovered proteases. In this screening, PrsW was not detected for some unknown reason, although σW activity was found to be impaired in a strain harboring a mutation in the ecsA gene (Heinrich et al., 2008; K. A., unpublished results). The mutation in ecsA was also revealed to block the proteolysis of RsiW by RasP. The gene ecsA exists in a putative operon of three genes, ecsA, ecsB and ecsC, which is predicted to encode a so-called ABC transporter complex, although the function of ecsC is still unknown. Inactivation of ecsA and/or ecsB (but not ecsC) in the operon caused a similar deficiency of SigW activity, suggesting that the transport ability of EcsA/EcsB was responsible for RasP regulation (Fig. 4). It was previously reported that ecs mutants showed pleiotropic phenotypes including decreased exoamylase (or possibly exoenzymes) and deficiencies of competence and sporulation in B. subtilis (the name of the operon is derived from these pleiotropic phenotypes) (Leskela et al., 1996). To date, the role of EcsABC, an ABC transporter involved in multiple events occurring after the logarithmic growth phase, in the regulation of RasP has not been elucidated precisely. One possible role is suggested from the experiment shown below. In the rsiW-deleted mutant, σW activity was higher than that in the wild type strain (Fig. 5A). Truncation of rsiW is also expected to cause loss of RsiW function, resulting in an increased level of σW activity. Unexpectedly, when the C-terminal, extracytosolic segment of RsiW was sequentially deleted, yielding ΔRsiW72 and ΔRsiW55, which still possessed 72 and 55 amino acids in the extracytosolic space (in contrast to 103 in wild type RsiW), the σW activity of cells decreased to levels lower than that of the wild type strain, indicating that the truncated RsiW strongly suppressed σW activity. On the other hand, further deletion, yielding ΔRsiW7, caused σW activity to increase, suggesting that ΔRsiW7 is unstable and susceptible to proteases. This observation suggested that the truncated RsiWs (ΔRsiW72 and ΔRsiW55) did not suffer further truncation with proteolysis, resulting in their being stabilized and rigidly binding to σW. It is proposed that the very end of the C-terminal region of RsiW would be required for interaction with PrsW, or that the peptide released from RsiW, cleaved at site 1 by PrsW, could induce further proteolysis at site 2 by RasP. Thus, in the latter case, the released peptide could be transported by or attached to EcsAB and this event would activate RasP (Fig. 4), although further analysis is still needed.
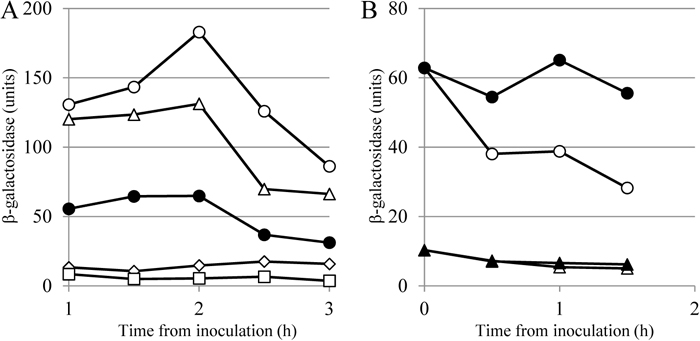
Altered expression of sigW under various conditions. Vertical axes indicate the β-galactosidase activities of the lacZ gene fused with the promoter sequence, which is recognized by σW. (A) Effect of RsiW deletion on its function of sequestering σW as an anti-sigma factor. Cells harboring the lacZ fusion and a progressively truncated series of RsiW were harvested at the indicated times and β-galactosidase activities were measured. Closed circles: rsiW+ (RsiW103); open circles: rsiW− (ΔRsiW0); open diamonds: ΔRsiW72; open squares: ΔRsiW55; and open triangles: ΔRsiW7. The superscript indicates the number of amino acids in the extracytoplasmic region, predicted by the PSORT program. (B) Effect of the addition of Zn ions on sigW expression. Cells of wild type (circles) and rasP mutant (triangles) in the absence (open symbols) and presence of 100 mM ZnCl2 (closed symbols) were harvested at the indicated times and the β-galactosidase activities were measured.
In S. coelicolor A3(2), the ECF sigma factor σR regulates the thioredoxin system in response to changes in the cytoplasmic thiol-disulfide status, and is controlled by an anti-sigma factor, RsrA, whose cytoplasmic (sigma factor binding) domain contains the CysXnHisX3CysX2Cys (CHCC) motif for the coordination of a zinc ion (Kang et al., 1999; Paget et al., 2001; Campbell et al., 2007). Under thiol-oxidizing conditions, RsrA forms intramolecular disulfide bonds and cannot bind σR, and thus discharges it. Therefore, this anti-sigma factor family is called the ZAS (zinc-binding anti-sigma factor) family. RsiW and YlaD (the putative anti-σYlaC) of B. subtilis have a CHCC motif in the N-terminal domain and are classified as ZAS family proteins (Paget et al., 2001). Although transcription of the ylaABCD operon depends on the ECF sigma factor σYlaC, and YlaD acts as an anti-sigma factor for σYlaC (Fig. 1), oxidative stress did not induce the transcription of ylaABCD (Matsumoto et al., 2005). Moreover, the interaction between σW and RsiW was not changed by oxidation of the CHCC motif in RsiW and the expression of sigW is unrelated to thiol-oxidizing conditions, suggesting that the RsiW activity for σW suppression is not regulated by the state of zinc coordination (Devkota et al., 2017). In B. subtilis, these CDCC motifs may have become silenced during the process of evolution, one (in YlaD) by changing into a pseudomotif and the other (in RsiW) by acquisition of additional mechanisms. According to our unpublished results, expression of sigW was higher in the presence of 100 μM ZnCl2 in the medium than in the absence of excess Zn ions (Fig. 5B). However, this elevation was not observed in the rasP mutant, suggesting that proteolysis of RsiW by proteases is regulated by zinc availability (possibly excess zinc stress).
The anti-σV, RsiV, is controlled by RIP, in which RasP also cleaves RsiV at site 2, whereas cleavage of RsiV at site 1 is performed by an unknown protease other than PrsW. Exposure of cells to the inducer lysozyme activates site 1 proteolysis, resulting in further truncation at site 2 by RasP (Hastie et al., 2013). It has been reported that E. coli outer membrane proteins unfolded by extracellular stresses bind to DegS, thus allowing DegS to cleave RseA (anti-σE) (Walsh et al., 2003). The molecular mechanisms that activate these controlling proteins are not yet understood precisely in B. subtilis, with the exception that the anti-σW factor RsiW is known to be regulated by multiple proteins in response to extracellular stresses. Since the structure of the cell surface in Gram-positive B. subtilis is distinct from that in Gram-negative E. coli, the underlying mechanisms are likely to be different. Expression of sigW is induced by various extracellular stresses including alkaline stress, cell wall-active antibiotics, antimicrobial peptides produced by other bacteria, and especially membrane stresses (Ho and Ellermeier, 2012; Helmann, 2016). The involvement of these different kinds of extracellular stresses in σW activation might explain the complexity of the regulatory proteins related to RsiW. The fact that almost all factors, like RsiW, PrsW, RasP and EcsAB, that are involved in the mechanism are membrane proteins raises the possibility that membrane destabilization, which is caused by the σW-activating stresses, leads to activation of these factors and then triggers the cleavage of RsiW by PrsW.
Regulation of σM in response to extracellular cell wall stressesσM is another well-studied ECF sigma factor in B. subtilis. The sigM operon is composed of three genes, sigM, yhdL and yhdK, and the latter two encode the putative membrane proteins YhdL and YhdK (Fig. 1). YhdL acts as the anti-σM and YhdK is predicted to interact only with YhdL and not with σM (Yoshimura et al., 2004). Abnormal overexpression of sigM caused by inactivation of yhdL provokes inhibition of cell growth (Horsburgh and Moir, 1999). Additionally, inactivation of yhdK causes retardation of cell growth, which is milder than that induced by yhdL inactivation, suggesting that YhdK binds to the σM-YhdL complex and supports YhdL function, e.g., by stabilization of the complex. The operon structure of sigM and yhdL is conserved in several bacterial species belonging to Firmicutes, but in some species yhdK is missing, indicating an auxiliary function for YhdK. It has been reported that more than 60 genes are regulated by σM (Asai et al., 2003; Eiamphungporn and Helmann, 2008). Notably, these include genes encoding the central components of cell division (e.g., div genes, min genes) and cell wall biosynthesis (e.g., mur genes, mre genes), which are essential for cell growth. This is consistent with the idea that disordered expression of these essential genes along with elevated expression of σM leads to lethality of the cell.
Expression of the sigM operon is upregulated in response to high salt, cell wall-active antibiotics, ethanol, heat, acidic pH and superoxide stress (Thackray and Moir, 2003). These stressors seem to affect cell wall integrity or function, whereas the stress signals against σW, including alkaline pH, seem to impair cell membrane stability. This discrimination is partly proven by the experiment shown in Fig. 6. The expression of sigM and sigW was induced when cells were exposed to acidic pH (6.0) and alkaline pH (8.0), respectively. The sigW expression induced by pH upshift was repressed by pH downshift, and the inversely repressed expression of sigW by pH downshift was immediately induced by pH upshift. In contrast, sigM expression induced by pH downshift was slightly repressed by pH upshift even though sigM expression repressed by pH upshift was not reversed by pH downshift. This discrepancy between σM and σW may reflect a difference in their mechanisms of regulation by the anti-sigma factor. In any case, these reversible and contrasting effects of pH shift on their expression may reflect a difference in a cell structure retaining the machinery that senses a stress signal between the two factors; namely, the structure is the cell wall for σM and the cell membrane for σW.
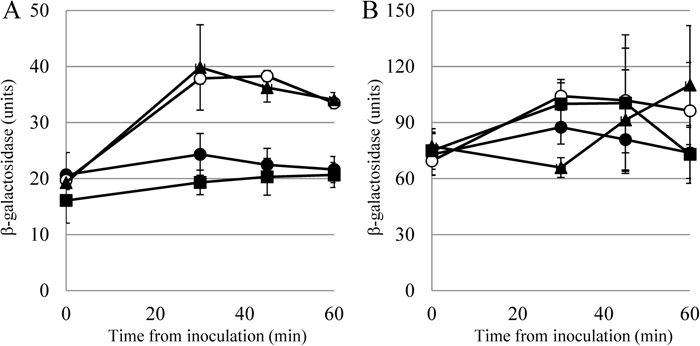
Effect of pH shift on the expression of sigM and sigW. The vertical axes indicate the β-galactosidase activities of the lacZ gene fused with the promoter sequence recognized by σM (A) and σW (B). Cells harboring the lacZ fusion were cultured in LB medium at various pHs, harvested at the indicated times, and the β-galactosidase activities were measured. Closed circles: pH 7.0 (normal condition); open circles: pH 6.0 (A) and pH 8.0 (B) (induction condition); closed squares: pH 8.0 to pH 6.0 after 30 min; closed triangles: pH 6.0 to pH 8.0 after 30 min. The experiments were performed repeatedly and the standard errors are indicated.
The overall processes of peptidoglycan synthesis are illustrated in the left part of Fig. 7. The basic units of a cell wall (N-acetylmuramic acid, N-acetylglucosamine and pentapeptide) are formed in the cytosolic space and assembled onto undecaprenyl phosphate, forming the so-called lipid II. Lipid II is then flipped to the outer surface of the cell membrane. Lipid II polymerization into a nascent peptidoglycan strand occurs via a glycosyltransferase-mediated peptidoglycan polymerization reaction. Cross-linking of peptide units in adjacent glycan strands occurs via a transpeptidase-mediated action (Lovering et al., 2012). Several steps are specifically inhibited by cell wall-active antibiotics. For example, fosfomycin blocks construction of the basic unit of the cell wall at the very first stage, moenomycin inhibits glycosyltransferase, vancomycin inhibits transpeptidase and bacitracin blocks the dephosphorylation of undecaprenyl diphosphate, resulting in defective recycling of undecaprenyl phosphate (Fig. 7). Such cell wall-active antibiotics are known to induce σM and several other genes, which catalyze steps of peptidoglycan synthesis and are included in the σM regulon. One of the σM regulon genes, bcrC (formerly ywoA), encodes a predicted undecaprenyl-diphosphate phosphatase (the target of bacitracin) (Ohki et al., 2003; Bernard et al., 2005). The cells of sigM-knockout mutants bulge out at the division site, which is indicative of cell wall defects (Horsburgh and Moir, 1999). As mentioned above, these observations strongly indicate that σM contributes to cell wall homeostasis against extracellular stress.
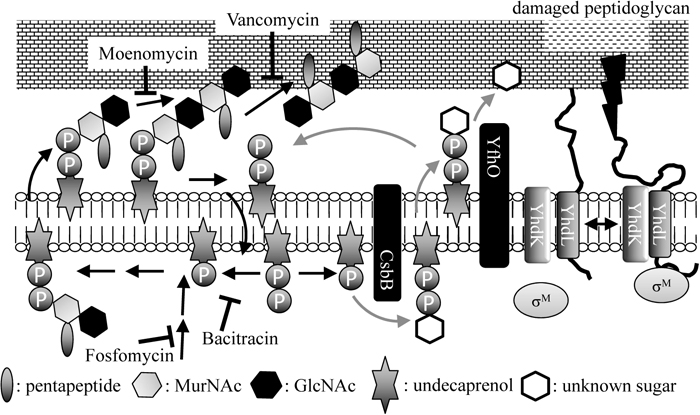
Diagram of peptidoglycan synthesis (left) and the regulation of σM by YhdL/K (right) in B. subtilis. The details are described in the text. The inhibition sites of some cell wall antibiotics are indicated by flat arrows. GlcNAc: N-acetylglucosamine; MurNAc: N-acetylmuramic acid.
To discover novel factors that influence the regulation of σM activity, transposon-mediated random mutagenesis was employed as in the case of σW. It was found that when yfhO, encoding a presumed bactoprenol-linked sugar translocase, is disrupted, expression of CsbB, which was already known to be transcribed with σB- (Table 1) and σX-containing RNA polymerases in response to extracellular stress (Akbar and Price, 1996; Huang and Helmann, 1998) and encodes a putative undecaprenol glycosyltransferase, causes constitutive activation of σM (Inoue et al., 2013). As shown in the right part of Fig. 7, it is proposed that accumulation of glycosylated undecaprenol diphosphate catalyzed by CsbB reduces the undecaprenol pool unless the undecaprenol is recycled after consumption of the sugar loaded on it by the action of YfhO. Shortage of available undecaprenol within a cell induces σM activity. The well-characterized system known as Gtr, including undecaprenol glucosyltransferases, determines serotype conversion (O-antigen glucosylation) in Gram-positive enteric bacteria (Korres et al., 2005). In B. subtilis, there are two CsbB homologs, YkcC and YkoT, in which essential amino acid residues residing in the catalytic domain are also conserved. The genes encoding YkcC and YkoT are preceded by genes encoding YkcB and YkoS, which possess a putative transmembrane region (14 and eight passes, respectively). Transcription of ykcBC is regulated by two-component regulatory systems of unknown function, YrkQ/P and YclJ/K (Ogura et al., 2008, 2010). YkcB and YkoS have no significant similarity in amino acid sequence with YfhO, which possesses 13 transmembrane domains. Nevertheless, similar to the case of CsbB and YfhO, deletion of ykcB and induction of ykcC leads to σM activation, whereas deletion of ykoS and induction of ykoT does not (K. A., unpublished results). In Gram-positive bacteria, peptidoglycan is further modified with carbohydrate-based anionic polymers such as lipoteichoic acid and wall teichoic acid, which plays an important role in cell envelope integrity. These polymers are decorated with sugar and phosphate, which may be modified after presentation on the cell surface by this presumed Gtr-like system in B. subtilis.
Experiments testing whether already-known proteases that decompose the anti-sigma factors are involved in proteolysis of YhdL and YhdK did not show a definitive result. Our preliminary data suggested that the intracellular concentration of epitope-tagged YhdL was unchanged even when the cells were exposed to σM-inducing stressors such as high salt. As shown in Fig. 6, σW activity changed instantly, increasing and decreasing with pH exchange, indicating nimble turnover of RsiW via proteolysis; in contrast, σM activity could not respond immediately to pH shift, suggesting the rather stable behavior of YhdL. These observations suggest that a conformational change in YhdL structure would be caused by extracellular stresses, resulting in release of σM (Fig. 7). Even so, it remains unknown whether direct sensing of cell wall damage by the C-terminal extension of YhdL protruding from the cell membrane (and possibly sensing the availability or abundance of undecaprenol or the lipid II intermediate via the transmembrane region of YhdL), and/or cell membrane degeneration caused by cell wall impairment, would lead to a conformational change in YhdL structure.
Universal effects of membrane degeneration on anti-sigma factorsThe molecular mechanisms regulating ECF sigma factors other than σW, σV and σM in B. subtilis have not been elucidated, although a few reports indicate that a broad effect on the composition and status of the cell membrane leads to global activation of ECF sigma factors. The processes of cell membrane biosynthesis, including lipoteichoic acid and glucolipids and their relationship with ECF sigma factors, are explained precisely in another review in this issue. Induction of σM and σV activities is observed when the lipid composition of membranes is altered by pgsA mutation, which causes a drastic reduction in phosphatidylglycerol content, whereas the others are not involved (Hashimoto et al., 2009). This demonstrates a specific relationship between the alteration of lipid composition in membranes and anti-sigma factor function. Lipoteichoic acid and glucolipids in the cell membrane are also required for maintenance of cell surface structure, and all the genes encoding ECF sigma factors except sigZ are induced in lipoteichoic acid synthesis-deficient cells (Hashimoto et al., 2013), while σM, σV and σX were found to be activated in glucolipid-lacking cells (Matsuoka et al., 2011). In a mutant strain with a disrupted shaA gene, which encodes the sodium-hydrogen antiporter, expression of σM and σW increased whereas that of σX was reduced. It was postulated that disruption of shaA damaged the proton gradient across the membrane, resulting in superoxide generation and cell membrane damage (Kosono et al., 2004). A counterpart of σW-RsiW exists in O. iheyensis, OIσW-OIRsiW. Heterologous expression of OIσW-OIRsiW in B. subtilis has a lethal effect on the cell, suggesting OIRsiW dysfunction and toxicity of the abnormally expressed OIσW (note that overexpression of σW is not toxic). The function of OIRsiW is restored by extension of the alpha helix in the transmembrane segment, suggesting that proper thickness of the cell membrane is required for the translocation and/or function of membrane proteins like RsiW (Yano et al., 2011). The transcription of fabF, which resides in an operon for fatty acid synthesis comprising fabHa and fabF, is regulated by σW through a promoter that exists within the fabHa gene (Kingston et al., 2011). Thus, upregulation of FabF via transcription induced with this promoter increases the average fatty acid chain length, resulting in a reduction of membrane fluidity and an increase of resistance to stress response. These observations indicate that the activity of ECF sigma factors is sometimes regulated by an anti-sigma factor via the universal effects of membrane degeneration; it is thus plausible that multiple stress response systems should be launched rapidly and simultaneously in order to confront an immediate and diverse environmental and internal exchange. Notably, glucose addition leads to increased levels of acetylated CshA, which is known to associate with RNA polymerase, and RNA polymerase bound to acetylated CshA could preferentially bind σX and σM (Ogura and Asai, 2016). This is a unique model of a regulatory link between ECF sigma factors and intracellular nutritional signals as well as extracellular stress signals, independent of anti-sigma factors.
Concluding remarksThe subfamily of ECF sigma factors was discovered recently after the emergence of whole-genome sequencing technology, in contrast to the gradual accumulation of studies about other sigma factors, even though the ECF sigma factors comprise a large proportion of all sigma factors found to date. This implies a particular physiological feature of ECF sigma factors; that is, these sigma factors are activated only under certain conditions, which rarely occur in laboratory conditions, and their functions are redundant in a single microorganism. Therefore, it is difficult to unearth these genes by means of typical genetic analyses because a representative phenotype emerging from the inactivation of an ECF sigma factor cannot easily be determined.
ECF sigma factors constitute the largest and most diverse subfamily within the sigma factor family. On average, most eubacteria possess a dozen ECF sigma factors. Protein subfamilies such as ECF sigma factors are predicted to be derived from a primary (housekeeping and essential) sigma factor and to have diverged by duplication and accumulation of mutations (Wösten, 1998). Comprehensive analysis of the phylogenetic relationship of ECF sigma factors was performed by comparative genomic prediction to seek an insight into their evolutionary history, and suggested that the signal transduction system with which the ECF sigma factors are involved is far more complex and diverse than had been previously observed and postulated (Staroń et al., 2009). Some ECF sigma factors are widely distributed in diverse bacterial phyla, indicating that they appeared before the individual phyla separated. However, the majority of ECF sigma factors are phylum-specific, implying that they have evolved individually to allow the species to specifically adapt to their environmental or cellular conditions.
As emphasized in this issue, diversity, hierarchy, specificity, complexity, and many other keywords of scientific interest in the field of molecular evolution and biology are included within the regulatory systems of transcription by ECF sigma factors, such as robustness acquired during the process of adaptation, interaction between ECF sigma factors and their cognate anti-sigma factors, and the molecular mechanisms of responses to the external environment via cell surface structures. Quests for answering how ECF sigma factors emerged and how they work will contribute to a comprehensive understanding of the biological regulatory network of gene expression.
I thank Fujio Kawamura, Naotake Ogasawara and Yoshito Sadaie for helpful discussions. This work was supported by JSPS Grants-in-Aid for Scientific Research (24580101 and 15K07350), and a JSPS Grant-in-Aid for Scientific Research on Innovative Areas (24119502).