2026 Volume 95 Issue 2 Pages 164-178
2026 Volume 95 Issue 2 Pages 164-178
Vernalization is a process in which plants achieve the competence to flower in the upcoming spring through exposure to long-term cold like winter. Unlike cold acclimation (short-term cold stimulus), vernalization is not immediately triggered by exposure to temporary low temperature. It seems likely that vernalization is triggered by low temperatures during winter that set up stable changes that last until the upcoming spring when they trigger floral transition. Over several decades, extensive studies have been conducted to elucidate the molecular mechanisms underlying the vernalization-mediated floral transition process using Arabidopsis thaliana (Arabidopsis) model plants, as well as crop plants. Comprehensive knowledge of the molecular mechanisms underlying vernalization-mediated floral transitions in the model plant Arabidopsis has helped us explore and reveal the molecular mechanisms controlling floral transitions in other plants, especially Brassica crops. This review highlights the current understanding of the molecular characteristics underlying vernalization-mediated flowering in Arabidopsis, as well as in several horticultural Brassica crop plants.
In temperate climates, winter-annual, biennial, and perennial plants sense long-term cold like winter and use it as an environmental signal for a suitable decision to time flowering in a favored season like spring (Bernier and Perilleux, 2005; Chouard, 1960). This requirement for long-term cold for flowering is known as vernalization. Vernalization serves to block floral transition in the fall season prior to winter, and then permits floral transition in the following spring season. Exposure to long-term cold stably switches the life cycle of plants from the vegetative to reproductive stage, particularly by suppressing a potent floral repressor gene, FLOWERING LOCUS C (FLC) (Amasino, 2010; Sheldon et al., 1999). Thus, vernalization is often referred to as an epigenetic phenomenon because a regulatory “switch” triggers the transcriptional modulation of FLC. This biological process is well aligned with the definition of epigenetic regulation. For instance, once FLC is repressed by vernalization, its silenced state is stably inherited by daughter cells even after the plants return to warm growth conditions. This long-term maintenance of gene expression status without changes to the underlying DNA sequence is one of the defining feature of epigenetics (Angel et al., 2011; Yang et al., 2014a). Vernalization-mediated silencing of FLC is mediated by chromatin modifications, particularly deposition of H3K27me3 by the Polycomb Repressive Complex 2 (PRC2) and removal of active histone marks like H3K4me3 and H3K36me3 on FLC chromatin. Involvement of chromatin-based gene regulation is also well aligned with the core principle of epigenetic gene regulation. Molecular mechanisms underlying vernalization-mediated epigenetic changes in plants have been intensively studied using Arabidopsis as a model system. In this review, we summarize current understanding of the molecular basis underlying vernalization-mediated floral transition. In addition, genes involved in Arabidopsis epigenetic mechanisms and several Brassica crops are described.
Vernalization is known to be perceived at the shoot apical meristem (SAM) in the shoot apex located at the tops of plants (Kim et al., 2009). The SAM consists of a population of pluripotent stem cells that produce organs like leaves and flowers in plants. Before floral transition occurs, SAM cells in plants maintain vegetative growth, generating leaves. The SAM directly senses cold and becomes vernalized. Once floral transition is initiated in a favored condition, the cell identity of a majority of SAM cells switch to the FAM (floral apical meristem), which generates flower organs like sepals, petals, stamens, and carpels. This was demonstrated by grafting experiments of plants exposed to different day lengths. Prior to flowering, the cells in the SAM must first become competent to respond to flowering signals (McDaniel et al., 1992).
Arabidopsis accessions can be divided into two groups depending on the vernalization requirement; vernalization requiring winter-annual versus summer-annual (early-flowering) accessions which do not require vernalization for flowering. For instance, winter-annual type Arabidopsis plants exhibit a delayed flowering phenotype compared to summer-annual plants. Intensive studies in the last few decades showed that the growth features of winter- and summer-annual plants is typically determined by the transcript level of FLC expression. Intensive genetic analyses using natural accessions between summer and winter types identified two major genes, FRIGIDA (FRI) and FLC, that confer a vernalization requirement in Arabidopsis (Clarke and Dean, 1994; Johanson et al., 2000; Koornneef et al., 1994). FRIGIDA (FRI) encodes a coiled-coil protein and was shown to interact with CBP20, a mRNA cap-binding protein, thus enhancing expression of FLC in some winter-type Arabidopsis accessions (Geraldo et al., 2009; Johanson et al., 2000). FLC encodes a MADS-box domain DNA binding protein that acts as a repressor of floral transition (Michaels and Amasino, 1999; Sheldon et al., 1999). A potent floral repressor, FLC directly binds to promoter regions of several key floral activators like FT (FLOWRING LOCUS T), FD, and SOC1 (SUPPRESSOR OF OVEREXPRESSION OF CO 1) to repress their expressions before vernalization (Searle et al., 2006). When expression of FLC is stably decreased by vernalization (winter season), floral activator genes like FT and SOC1 can be greatly induced by the CO (CONSTANS)-mediated photoperiod flowering pathway in the following spring season (Golembeski et al., 2014). Subsequently, FT and SOC1 stimulate other floral identity genes such as AP1 (APETALA 1) (Abe et al., 2005; Lee and Lee, 2010). Activated AP1 protein interacts with its cofactors like SEP3 (SEPELLATA 3) and LFY (LEAFY) to form floral organs in the FAM by positively regulating AG (AGAMOUS), AP3 (APETALA 3), and PI (PISTILLATA) (Kaufmann et al., 2009).
FLC is initially activated during embryogenesis in Arabidopsis. A study reported that activation of FLC during embryogenesis is initiated by a NF-YB (NUCLEAR FACTOR Y Subunit B) type transcription factor, designated LEAFY COTYLEDON 1 (LEC1) (Tao et al., 2017). LEC1 directly recognizes and binds to the CCAAT motif within the promoter region of FLC during early embryogenesis and coordinates the accumulation of active histone marks like H3K4me3 and H3K36me3 in the FLC chromatin. In addition, LEC1 physically associates with seed-specific B3 domain transcription factors like LEC2 (LEAFY COTYLEDON 2), ABSCISIC ACID-INSENSITIVE 3 (ABI3), and FUSCA3 (FUS3) during embryogenesis (Santos-Mendoza et al., 2008; Tao et al., 2019; Xu et al., 2022). Interestingly, the B3 domain transcription factors, LEC2, ABI3, and FUS3 specifically bind to the RY motif (-CATGCA-) located in the 1st intron region of genomic FLC (Tao et al., 2019; Xu et al., 2022). Thus, it is proposed that a pioneer protein, LEC1 binds to the CCAAT motif located at the distal promoter region of FLC, promoting the recruitment of LEC2, ABI3, and FUS3 to the RY motif in the 1st intron of FLC for the initiation of FLC expression. Taken together, the coordinated action of LEC1-LEC2-ABI3-FUS3 on the FLC genomic region may trigger the expression of FLC during embryogenesis in Arabidopsis.
After initial activation of FLC during embryogenesis, LEC2 and FUS3 were shown to associate with components of the FRI-C activation complex, catalyzing H3K4me3 and H3K36me3 on FLC chromatin. The transcription factors associated with the FRI-C protein complex accelerate the stable establishment of an active chromatin environment around the FLC region (Li et al., 2018). High expression of FLC during the early growth period is substantially correlated with inhibition of floral transition in Arabidopsis plants. Intensive genetic screening identified components of activation complexes conferring high FLC expression (Choi et al., 2007, 2011; He et al., 2004). For instance, FRI acts as a scaffold protein interacting with FRIGIDIA-LIKE 1 (FRL1), FRIGIDA ESSENTIAL 1 (FES1), SUPPRESSOR OF FRI 4 (SUF4), and FLC EXPRESSOR (FLX) to form a transcription activator complex and activate expression of FLC (Choi et al., 2011). The FRI-complex probably recruits another chromatin complex, the SWR1-complex. Some FRI-complex components physically associate with components of the SWR1-complex such as PIE1 (PHOTOPERIOD-INDEPENDENT EARLY FLOWERING 1), SWC6/SEF (SERRATED LEAVES AND EARLY FLOWERING), YAF9/TAF14 (TBP-ASSOCIATED FACTOR 14), SUF3 (SUPPRESSOR OF FRI 3)/ARP6 (ACTIN-RELATED PROTEIN 6), and ARP4 (ACTIN-RELATED PROTEIN 4). The SWR1-complex is involved in ATP-dependent chromatin remodeling, particularly by replacing a histone variant, H2A.Z, in the target chromatin to confer transcriptional flexibility in response to endogenous and environmental changes (Deal et al., 2007; March-Diaz and Reyes, 2009). Therefore, physical interaction between FRI-complex components and the SWR1-complex complexes suggests that FLC chromatin undergoes dynamic changes in its chromatin context.
The PAF1 (POLYMERASE II-ASSOCIATED FACTOR 1)-containing complex is also required for the activation of FLC via stimulating accumulation of H3K4me3 on FLC chromatin (He et al., 2004; Oh et al., 2004; Yu and Michaels, 2010; Zhang and van Nocker, 2002). The Arabidopsis PAF1-complex includes ELF7 (EARLY FLOWERING 7), ELF8 (EARLEY FLOWRING 8), VIP3 (VERNALIZATION INDEPENDENCE 3), VIP4 (VERNALIZATION INDEPENDENCE 4), VIP5 (VERNALIZATION INDEPENDENCE 5), and PHP (PLANT HOMOLOGOUS TO PARAFIBROMIN) components. Mutants of these PAF1-complex components displayed a rapid flowering phenotype compared to wild type plants, demonstrating that they are commonly required for activation of FLC in Arabidopsis. Another epigenetic regulator, EFS (EARLY FLOWERING IN SHORT DAYS)/SDG8 (SET DOMAIN GROUP 8) interacts with the FRI-complex and catalyzes H3K4me3 and/or H3K36me3, which are histone marks for FLC activation (Kim et al., 2005; Ko et al., 2010; Li et al., 2015; Xu et al., 2008). EFS protein is highly enriched at FLC chromatin before vernalization. As a result, efs mutants displayed a rapid floral transition due to low expression of FLC. Taken together, the FRI-complex plays a critical role in recruiting and facilitating the catalytic activity of the SWR1-complex and PAF1-complex complexes to activate FLC.
In Arabidopsis accessions, winter-annual accessions exhibit significantly delayed flowering behavior compared to that of summer-annual accessions if they were not exposed to vernalization. Genetic approaches using winter-annual plants were conducted to isolate some mutants showing altered vernalization response. Several genes including VERNALIZATION 2 (VRN2), VERNALIZATION 1 (VRN1), VERNALIZATION INSENSITIVE 3 (VIN3), VERNALIZATION 5 (VRN5)/VIN3-LIKE 1 (VIL1), and AtPRMT5A (ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 5A) were identified (Gendall et al., 2001; Greb et al., 2007; Levy et al., 2002; Sung and Amasino, 2005). Loss-of-function mutants of these genes failed to stably maintain an FLC repressed state in response to vernalization. Among these factors, expression of VIN3 displayed an interesting pattern. For instance, VIN3 is only substantially induced by exposure to long-term cold and is completely abolished when plants return to warm temperatures (Sung and Amasino, 2004b). The inductive and transient nature of VIN3 expression suggests that it may play a key role in triggering subsequent molecular events to suppress FLC while plants are in vernalizing cold conditions. A recent study showed that two circadian regulators, CCA1 (CIRCADIAN CLOCK ASSOCIATED 1) and LHY (LATE ELONGATED HYPOCOTYL) redundantly act to activate expression of VIN3 during vernalization through the recognition of a vernalization-responsive cis-element (named VREVIN3) located in the proximal promoter region of VIN3 (Kyung et al., 2022). In a cca1;lhy double mutant with a winter-annual plant background, induction of VIN3 was significantly impaired under long-term cold, especially under mild cold conditions such as at 12°C. The VREVIN3 element contains G-box and EE (evening element) sequences. It was demonstrated that mutations in either the G-box or EE compromised full induction of VIN3 upon vernalization, leading to defects in the vernalization response. However, this study found that another unknown sensory module, which functions in concert with these circadian regulators, is needed for full induction of VIN3 in response to vernalization.
Epigenetic suppression of FLC chromatin by vernalizationGenetic isolation of VIN3, VIL1, VRN2, and AtPRMT5A strongly suggested the possible involvement of chromatin level regulation of FLC because all of these genes were reported in histone modification in animals or Drosophila. For instance, VRN2 encodes a homolog of Suppressor of zeste (Su(z)12), which is one of core components of Polycomb Repression Complex 2 (PRC2) for Histone H3 Lys 27 (H3K27) methylation in animals (Gendall et al., 2001). VIN3 and VIL1 commonly encode the PHD (Plant Homeo Domain) finger protein which was reported to act as a chromatin reader involved in chromatin level regulation (Franco-Echevarria et al., 2023). AtPRMT5A is involved in epigenetic regulation by methylating histones, particularly H4R3me2s (symmetric dimethylation of histone H4 at arginine 3) (Schmitz et al., 2008).
When examining the histone composition in the FLC gene region, two repressive histone modifications including Histone H3 Lys 9 (H3K9) and H3K27me2/3 methylation were greatly increased in FLC chromatin (Bastow et al., 2004). Meanwhile, in the vin3 mutant, these repressive histone marks such as H3K9me2 and H3K27me2/3 were not enriched in the FLC region. This led to the failure of FLC suppression, resulting in significantly delayed flowering. It was also demonstrated that these repressive histone marks were not increased in FLC chromatin in the vrn2 mutants (Gendall et al., 2001). VIN3 and VIL1/VRN5 belong to a subgroup of PHD finger domain proteins. VIL1/VRN5 was identified by yeast two hybrid screening for VIN3 interacting proteins (Sung et al., 2006) and genetic screening for mutants with altered vernalization responses in winter-annual Arabidopsis plants (Greb et al., 2007). Similar to VIN3, VIL1/VRN5 was also needed for the enrichment of H3K9 and H3K27 methylation in FLC chromatin by vernalization.
Several studies suggested that PHD domains can bind to various histone tails, ranging from non-methylated histone H3 (Lan et al., 2007), methylated histone H3 Lys 9 (Karagianni et al., 2008), and methylated histone H3 Lys 36 (Shi et al., 2007). These diverse binding activities of PHD finger domains make them potential candidates as “histone readers” of histone modifications. Amino acid sequence variations within PHD finger domains may be responsible for such variations in their specificity (Li et al., 2007). PHD finger domains in VIN3 family proteins contain common amino acids for the C3HC4 zinc finger; however, they also have variations in amino acid sequences including aromatic amino acids that create a binding pocket for the histone tail (Kim and Sung, 2013). A study showed that the specificity of VIN3/VIL1 PHD finger domains was responsible for the recognition of di-methylated histones H3 like H3K4me2 and H3K9me2 (Kim and Sung, 2017).
Role of the PRC2 complex in the suppression of FLC in response to vernalizationGenetic identification of VRN2, an Arabidopsis homolog of Su(z)12 raised the possibility that vernalization-mediated suppression of FLC might involve a complex similar to PRC2 in animals. PRC2 is a repressive chromatin regulator that acts to catalyze the methylation of H3K27 on target chromatins to suppress target gene transcription. Polycomb (PcG) group proteins were first identified from Drosophila as an essential component for maintenance of the repressive state in homeotic HOX genes (Schuettengruber et al., 2007; Schwartz et al., 2010). PcG proteins function as multi-protein complexes, known as Polycomb Repression Complexes (PRCs). These PRC complexes can be grouped into two distinct complexes, PRC2 and PRC1 (Polycomb Repression Complex 1). The PRC2 complex is responsible for the initial di- or tri-methylation of H3K27 at target gene chromatins. Methylated H3K27 can be recognized and bound by the PRC1 complex for subsequent chromatin compaction and stable gene suppression. Core components of PRC2 are well conserved throughout higher eukaryotes, including plants. In Drosophila, core components of PRC2 are made up of Enhancer of Zeste (E(z)), Extra sex combs (Esc), Suppressor of zeste Su(z)12, and P55 (Nurf55/RbAp48/CAF-1p55).
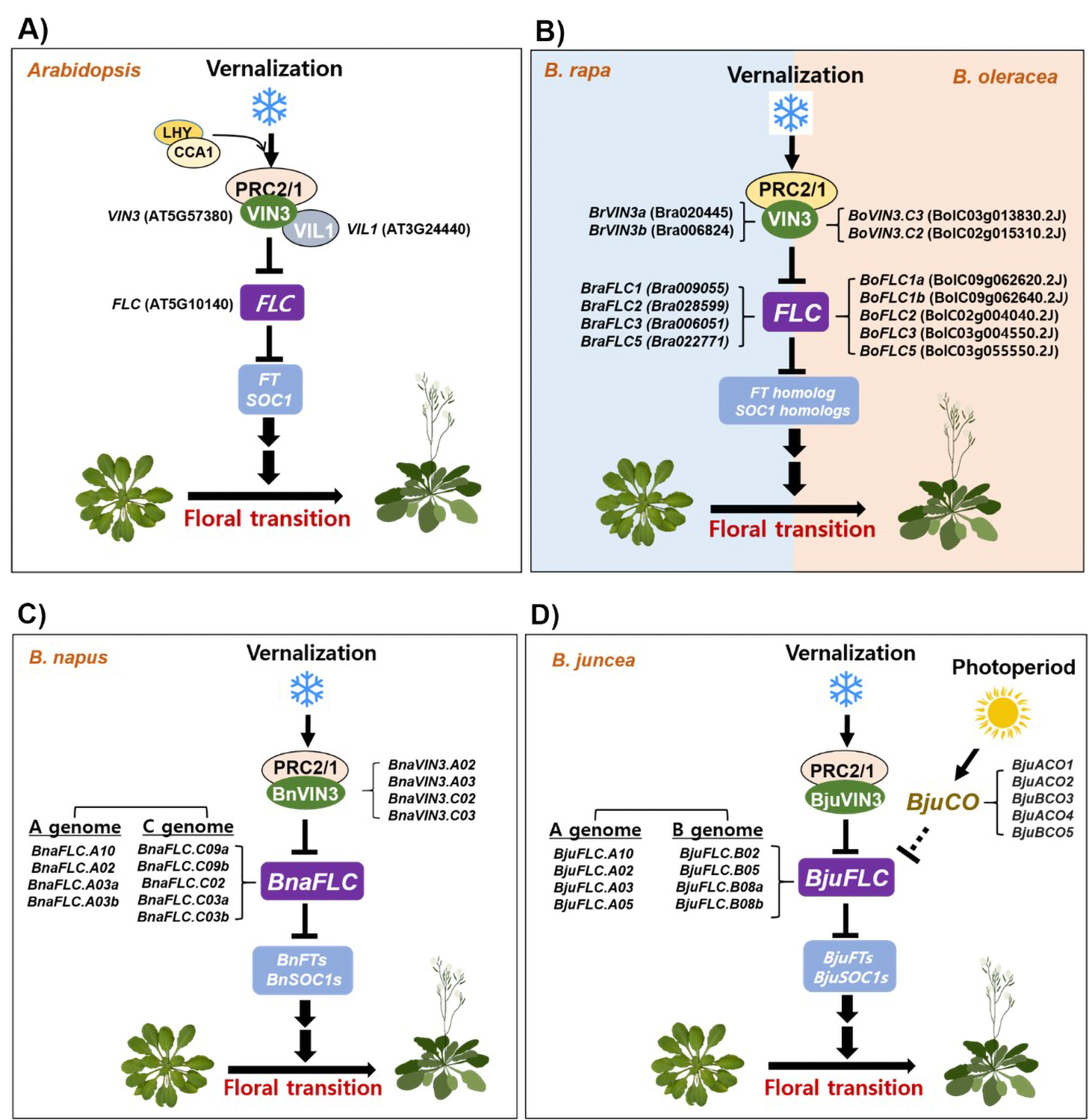
The Arabidopsis genome has PcG proteins similar to Drosophila PRC2 (Hennig and Derkacheva, 2009; Kim and Sung, 2014). In Arabidopsis, there are three homologs of Drosophila E(z)s, including CURLY LEAF (CLF), SWINGER (SWN) and MEDEA (MEA) (Grossniklaus et al., 1998). CLF was first identified as a repressor of the AG floral homeotic gene in Arabidopsis (Goodrich et al., 1997). SWN is the closest paralog of CLF and functions in part redundantly with CLF (Chanvivattana et al., 2004). MEA was identified from mutants with defects in endosperm development (Jullien et al., 2006; Kinoshita et al., 1999). Biochemical purification using VIL1/VRN5-TAP transgenic plants in vernalization showed that Arabidopsis PRC2 includes core PRC2 components (VRN2, SWINGER [an E(Z) homologue], FIE (FERTILIZATION-INDEPENDENT ENDOSPERM) [an ESC homolog], MSI1 (MULTICOPY SUPPRESSOR OF IRA1) [p55 homolog]), and PHD finger domain proteins such as VRN5/VIL1, VIN3, and VEL1 (VERNALIZATION 5/VIN3-LIKE 1)/VIL2 (VIN3-LIKE 2) (De Lucia et al., 2008; Wood et al., 2006). Therefore, the Arabidopsis genome contains highly conserved homologs of the Drosophila PRC2 complex, catalyzing tri-methylation of H3K27 at target gene chromatins including FLC. Stable maintenance of FLC repression following vernalization requires the PRC2 protein complex, which includes VIN3. This results in the de-repression of several floral integrator genes like FT and SOC1 which act as a floral activator triggering floral transition of Arabidopsis from the vegetative to the reproductive stage (Fig. 1A).
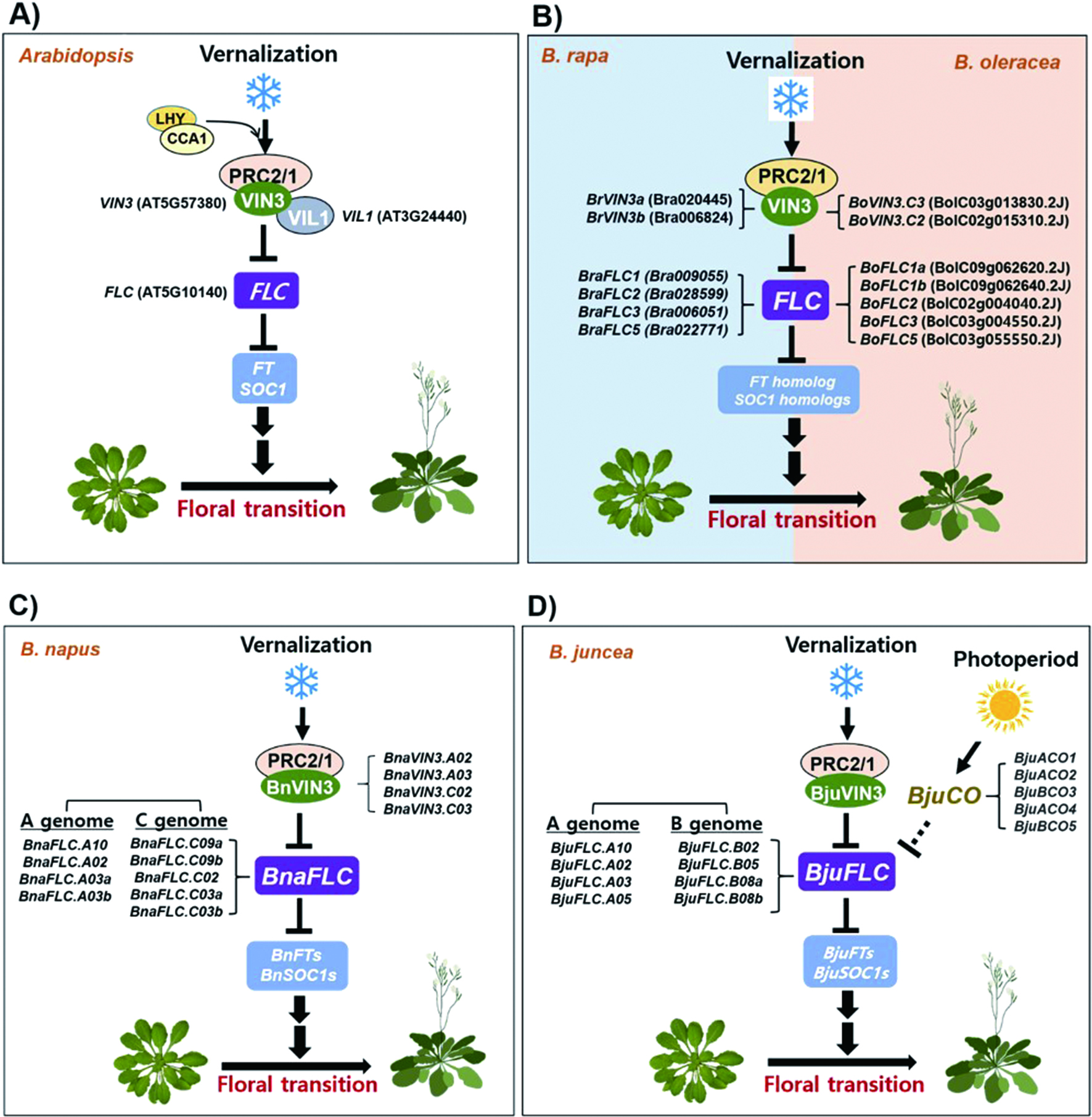
A schematic diagram showing the vernalization-mediated floral transition of Arabidopsis and Brassica genus plants.
Vernalization-mediated repression of FLC occurs in two steps: (1) accumulation of H3K27me3 at the FLC nucleation site during cold exposure, and (2) spreading of this mark across the entire FLC locus after the plant returns to warm temperatures (Yang et al., 2017). During vernalization, the PRC2 complex is enriched at the nucleation site, and upon return to warmth, H3K27me3 spreads throughout the FLC region during mitotic divisions, stably maintaining FLC repression. This process forms the basis of the “epigenetic memory of winter”. The PRC1 complex, including LIKE-HETEROCHROMATIN 1 (LHP1), reinforces this memory by recognizing H3K27me3 and promoting its further spread via positive feedback (Veluchamy et al., 2016; Yang et al., 2017).
Ground-breaking studies revealed that RY motifs (CATGCA/TGCATG) within the 1st intron of FLC help recruit the PRC2 complex during vernalization (Questa et al., 2016; Yuan et al., 2016). Disruption of RY motifs substantially failed to suppress expression of FLC, resulting in delayed floral transition. This DNA element is known as the Cold Memory Element (CME) (Yuan et al., 2016). In addition, B3 domain transcription factors, VP1/ABI3-LIKE 1 (VAL1) and VAL2 were demonstrated to recognize and bind to CME within the 1st intron of FLC (Questa et al., 2016; Yuan et al., 2016). Furthermore, VAL1 and VAL2 physically associate with MSI1, one component of the PRC2 complex and LHP1, one component of the PRC1 complex (Chen et al., 2018b, 2020; Yuan et al., 2016, 2021). Based on these findings, CME-binding VAL1/VAL2 is thought to recruit the MSI1-containing PRC2 complex and LHP1-containing PRC1 complex to enrich H3K27me3 on FLC chromatin upon vernalization.
Accumulated H3K27me3 at the FLC locus is maintained and inherited by daughter DNAs via a DNA-replication-coupled mechanism in Arabidopsis (Jiang and Berger, 2017). For instance, a histone chaperone complex named CHROMATIN ASSEMBLY FACTOR 1 (CAF1) incorporates a canonical H3.1-H4 histone tetramer into a newly generated daughter DNA strand during the DNA replication process, on which H3K27me3 is newly catalyzed on FLC chromatin (Jacob et al., 2014; Jiang and Berger, 2017; Yu et al., 2019). FASCIATA 1 (FAS1), one component of the CAF1 complex, physically associates with PCNA1 (PROLIFERATING CELL NUCLEAR ANTIGENS 1) and PCNA2, which are involved in the DNA replication process. In addition, it was demonstrated that FAS1 interacts with a couple of PRC complex components such as CLF, LHP1, and RING1A (RING FINGER PROTEIN 1) (Jiang and Berger, 2017). Therefore, CAF1 seems to incorporate the H3.1/H4 histone tetramer to newly synthesized histones during DNA replication. Concurrently, CAF1 helps to recruit PRC2/1 complexes which catalyze H3K27me3 to the newly incorporated H3.1/H4 histone tetramer for the stable transmission of parental epigenetic contexts to daughter DNAs.
Recently, a novel conceptual framework based on digital mathematical modeling has been proposed to explain the quantitative nature of vernalization-induced repression of FLC (Angel et al., 2011; Menon et al., 2021; Satake and Iwasa, 2012; Yang et al., 2017). For example, a study introduced a model in which the quantitative aspect of vernalization arises from H3K27me3-mediated silencing of FLC in a subset of cells. The proportion of cells in which FLC is silenced via H3K27me3 increases with the duration of cold exposure (Angel et al., 2011). Similarly, Satake and colleague developed a mathematical model that captures the dynamics of FLC expression, emphasizing its bistable nature, in which the gene exists in either an ON or OFF state (Satake and Iwasa, 2012). This model explains how environmental cues like temperature changes can affect the ON/OFF switch between these states. This provides a theoretical framework for understanding the memory aspect of vernalization. These studies collectively propose that FLC expression behaves in a binary fashion at the single-cell level, supporting the idea of a digital, switch-like mechanism in epigenetic memory. This binary expression pattern enhances the robustness of the vernalization response by ensuring clear, stable gene regulation at the cellular level. Moreover, such mathematical modeling helps unify mechanistic insights across different chromatin regulators, including those studied in mammalian systems (Bintu et al., 2016). This perspective has important implications for understanding how plants quantitatively measure cold duration and may help answer long-standing questions about the molecular mechanisms underlying environmental memory.
Brassica is a genus containing approximately 46 flowering plant species (Table 1) (Kiefer et al., 2014). The knowledge gained from Arabidopsis model plants has great potential to be extensively applied to Brassica genus crops to improve flowering time control and yield optimization. Thus, recently many studies have tried to elucidate the molecular details underlying the vernalization-mediated floral transition process in crops, especially Brassica plants. Here, the current understanding of molecular circuitries working in the vernalization response in several Brassica crop plants is briefly described.

List of 46 Brassica genus plants. This information was obtained from the BrassiBase website (https://brassibase.cos.uni-heidelberg.de/) and a reference (Kiefer et al., 2014).
Brassica rapa crops are important leafy vegetables cultivated worldwide. Flowering time is a critical agronomic trait in Brassica rapa leafy vegetables, as the commercial value of these crops depends on the optimal development of vegetative organs rather than reproductive structures (Itabashi et al., 2019; Shah et al., 2018). Premature flowering, or bolting, not only diverts energy away from leaf production but also leads to the formation of tougher, more fibrous, and often bitter leaves, significantly reducing both yield and marketability. Many Brassica rapa cultivars are highly sensitive to environmental cues such as temperature and photoperiod. Premature bolting triggered by suboptimal or fluctuating climatic conditions can substantially compromise agronomic performance and yield stability. Therefore, selecting or engineering cultivars with delayed or environmentally stable flowering behavior are essential for ensuring a longer vegetative growth phase, broader planting windows, and overall crop resilience. Improved control of flowering time can enhance productivity and adaptation in diverse growing regions, making it a key target in breeding programs for leafy Brassica crop plants (Bloomer and Dean, 2017). For these reasons, many studies have been conducted to elucidate the molecular mechanism underlying the environmentally-triggered floral transition (e.g., vernalization) in B. rapa crop plants. A study reported that several quantitative trait loci (QTL) affecting floral transition in B. rapa were identified using an F2 mapping population developed by crossing between annual and biennial lines (Osborn et al., 1997; Teutonico and Osborn, 1995). The reference genome of the B. rapa cultivar ‘Chiifu-401-42’ contains four FLC homologs known as BrFLC1 (Brassica rapa FLC 1) (Bra009055), BrFLC2 (Bra028599), BrFLC3 (Bra006051), and BrFLC5 (Bra022771) (Table 2). Several studies using an F2 mapping population reported that BrFLC1 and BrFLC2 were significantly linked to the QTLs controlling floral transition (Li et al., 2009; Wu et al., 2012; Xiao et al., 2013; Zhao et al., 2010). In a different F2 population (Early × Tsukena No. 2), BrFLC2 and BrFLC3 were co-localized with the QTL peaks, influencing floral transition in response to vernalization (Kakizaki et al., 2011). In addition, it was revealed that genomic sequences of BrFLC2 and BrFLC3 in the late-flowering line Tsukena No. 2 possessed large DNA insertions within their first intron region (Kitamoto et al., 2014). It was proposed that the sequence element in their first intron regions may play a critical role in the vernalization-mediated floral transition in B. rapa. Another study showed that over-expression of BrFLC1, BrFLC2, and BrFLC3 in Arabidopsis resulted in significantly delayed flowering time, indicating that BrFLC homologs commonly play a negative role in the floral transition in B. rapa, similar to that of Arabidopsis FLC (Huang et al., 2018; Kim et al., 2007).
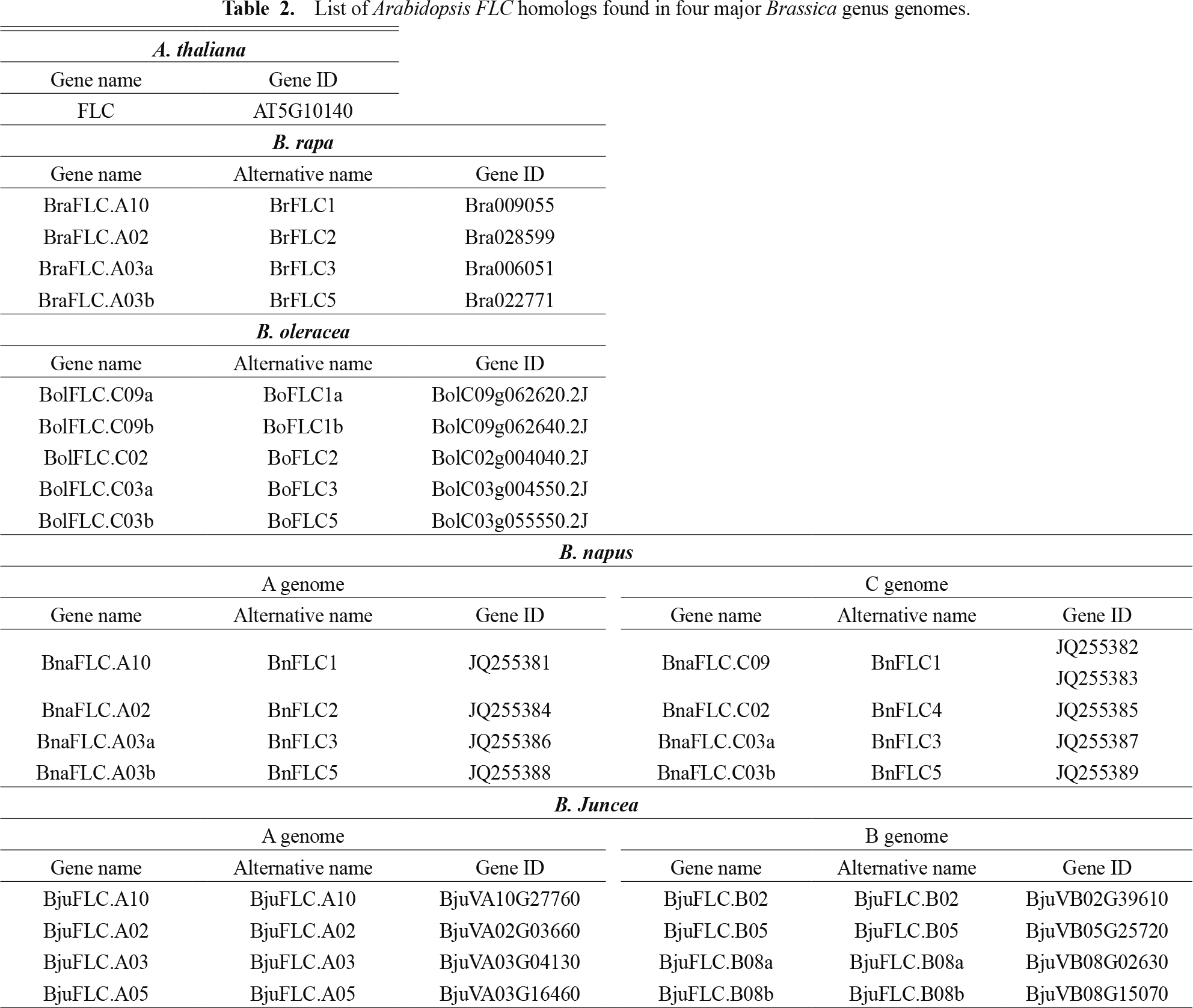
List of Arabidopsis FLC homologs found in four major Brassica genus genomes.
Expressions of all four BrFLC homologs were reduced by exposure to long-term cold and stably maintained at low levels even after plants were returned to warm growth temperatures (Kawanabe et al., 2016). Active histone marks like H3K4me3 and H3K36me3 were substantially accumulated at chromatins of BrFLC homologs prior to vernalization, promoting active expression of BrFLC homologs. However, when exposed to long-term cold, two B. rapa homologs of VIN3 were substantially induced by vernalization. During vernalization, induced BrVIN3 may cooperate with B. rapa PRC2 and (highly likely with) PRC1 complexes catalyzing accumulation of repressive histone marks like H3K27me3 on the BrFLCs regions. Even after plants return to warm growth temperatures, the repressed state of BrFLCs is stably maintained. Stable repression of BrFLCs results in the de-repression of floral integrators like two BrFT (Brassica rapa FLOWERING LOCUS T) homologs (BrFTa and BrFTb) and three BrSOC1 (Brassica rapa SUPPRESSOR OF OVEREXPRESSION OF CO 1) homologs (BrSOC1a, BrSOC1b, and BrSOC1c) which promote the floral transition of B. rapa (Kang et al., 2022; Kim et al., 2022). This indicates that the epigenetic histone modification context plays a crucial role in the transition from the vegetative to the reproductive stage by affecting transcription of the BrFLC genes in B. rapa plants, similar to Arabidopsis (Fig. 1B left).
Brassica oleraceaB. oleracea plants include many commercially popular vegetables and can be grouped according to their edible parts. For instance, cabbage (B. oleracea L. var. capitata), kale (Brassica oleracea var. sabellica), and kohlrabi (Brassica oleracea var. gongylodes) are harvested at the vegetative stage, while broccoli (B. oleracea L. var. italica) and cauliflower (B. oleracea L. var. botrytis) are harvested for their curd (edible flower head part of the plant) at the reproductive stage. Hence, understanding the molecular mechanism underlying the control of the flowering time of these Brassica species is of great importance.
Similar to B. rapa, four FLC homologs were identified from the B. oleracea genome (Table 2) and reported to be involved in the regulation of the B. oleracea flowering time (Leijten et al., 2018; Lin et al., 2005). By QTL analysis, BoFLC4 (also called as BoFLC2) was identified as a causative gene conferring the broccoli vernalization requirement (Irwin et al., 2016; Okazaki et al., 2007), as well as in cabbage (Okazaki et al., 2007), and cauliflower (Ridge et al., 2015). For instance, a study including biennial, annual, and rapid-cycling lines of cabbage plants showed that a frameshift mutation in exon 4 resulted in production of a BoFLC2 in-frame stop codon, leading to an early flowering phenotype (Okazaki et al., 2007). When the genomic sequence of BoFLC4 was introduced into the Arabidopsis flc-2FRI mutant, which carries a large genomic deletion at the FLC locus and exhibits an extremely early flowering phenotype (Michaels and Amasino, 1999), expression of BoFLC4 fully rescued the early flowering phenotype. This indicated that BoFLC4 plays a role as a floral repressor and contributes to conferring the vernalization requirement of B. oleracea plants (Irwin et al., 2016). In another QTL analysis, BoFLC3, BoFLC5, and BoFLC1 were also identified as a major locus co-localizing with QTLs responsible for flowering time regulation (Razi et al., 2008). Another study reported that the BoFLC3 locus is closely correlated with curd induction timing in subtropical broccoli breeding lines under a subtropical environment (Lin et al., 2018). A recent study showed that deletion of a 215-bp DNA fragment within the 1st intron region of BoFLC2 resulted in an impaired vernalization response, delaying cabbage flowering (Li et al., 2022). This is consistent with other cases reported in B. rapa and Arabidopsis, which showed that sequence variation within the 1st intron of FLC and BrFLC homologs resulted in variation in the vernalization response (Kitamoto et al., 2014; Questa et al., 2016; Yuan et al., 2016). In BoFLC1, two tandemly duplicated BoFLC1 homologs (designated BoFLC1a and BoFLC1b) were found in the cabbage genome database (Table 2). These genes were also demonstrated to play a crucial role in regulating flowering time in cabbage by acting as a floral repressor responsive to vernalization. For instance, in the non-flowering cabbage mutant line ‘nfc’, the duplicated BoFLC1a and BoFLC1b homologs maintained high expression levels throughout winter, in contrast to normal flowering lines in which BoFLC1 expression significantly declined (Kinoshita et al., 2023). Sustained expressions of BoFLC1a and BoFLC1b substantially suppressed floral integrator genes like BoFTs, inhibiting the transition to flowering even after vernalization. Its expression dynamics, influenced by genetic variations and duplications, determine the transition from the vegetative to reproductive stage of cabbage. Another study showed that a 67-base pair insertion in the second intron of the BoFLC1.C9 allele (BoFLC1) is correlated with delayed downregulation of BoFLC1 by vernalization, leading to later flowering (Abuyusuf et al., 2019). Expression analyses before and after vernalization at different plant ages revealed that three BoFLC homologs (BoFLC1~BoFLC3) responded differently to prolonged cold exposure. Regardless of plant age, BoFLC1 showed only a modest decrease, while BoFLC2 and BoFLC3 were strongly repressed by sufficiently long cold treatment (Itabashi et al., 2019). These differential expression patterns imply that each BoFLC homolog has a unique role in regulating flowering time in cabbage, with BoFLC2 and BoFLC3 being sensitively responsive to vernalization in cabbage. Collectively, these results indicated that BoFLC homologs are functionally well conserved and involved in the floral transition of B. oleracea in a similar way to Arabidopsis and B. rapa (Fig. 1B right). However, the detailed molecular mechanism underlying vernalization-mediated floral transition in B. oleracea is still largely unknown. For instance, it is still not understood as to why cabbage has four BoFLC homologs and whether or not they share functional redundancy and diversity. To address this point, development of knock-out mutants of BoFLC homologs using CRISPR/Cas9 genome editing methods may accelerate molecular studies to elucidate the detailed roles of each BoFLC member in the floral transition control of B. oleracea plants.
Brassica napusRapeseed (Brassica napus) is an allopolyploid plant popularly cultivated as a major oil crop that was created by hybridization between two diploid species, Brassica rapa (AA genome, 2n = 20) and Brassica oleracea (CC genome, 2n = 18). Thus, the B. napus genome contains both genomes (AACC, 2n = 38) (Chalhoub et al., 2014) (Table 2). Rapeseed can be classified into three groups based on the vernalization requirement: a winter type, a semi-winter type, and a spring type. Winter type B. napus plants are biennials and grow well in cooler temperatures and under high humidity in temperate climates. They have an obligate vernalization requirement to acquire competence for flowering. Semi-winter types are less dependent on vernalization for flowering and are grown in regions with moderate winter temperatures, such as central China. Meanwhile, spring type B. napus lines are annual plants which flower in the same year of sowing regardless of the presence of vernalization (Leijten et al., 2018).
In an Arabidopsis model plant, FRI was one of the major determinants of the vernalization requirement (Shindo et al., 2005). Four FRI homologs were identified in the genome of B. napus (Table 3). Among them, BnaA.FRI.a was co-localized with a major QTL peak for flowering time using a doubled-haploid population derived from a cross between the European winter cultivar ‘Tapidor’ and the Chinese semi-winter cultivar ‘Ningyou7’ (Wang et al., 2011). A transgenic analysis using functional complementation of BnaFRI.A03 under its native promoter exhibited a significantly delayed flowering of an Arabidopsis model plant (Yi et al., 2018). In agreement with delayed flowering, expression of Arabidopsis FLC was substantially higher in complemented lines with a BnaFRI.A03 transgene cloned from the winter type ‘Tapidor’ than that derived from the spring type ‘Ningyou-7’ cultivar. Analysis using promoter swap constructs suggested that genomic coding sequence variation within BnaFRI.A03 was critical for determining flowering time behavior. Meanwhile, expression variation did not affect the difference in flowering time. However, the causal SNPs have not yet been identified. In addition, Loss of BnaFRI had no significant effect on BnaFLC expression. The expression levels of BnaFLC genes were not reduced in the absence of BnaFRI, suggesting that additional loci independent of BnaFRI contribute to the low BnaFLC expression observed in spring type cultivars. This indicates that other genetic factors, such as variations in BnaFLC cis-regulatory elements, epigenetic modifications, or alternative repressors, may be responsible for the reduced BnaFLC expression, leading to an early-flowering phenotype in spring cultivars. This observation aligns with findings in Arabidopsis thaliana, where loss-of-function mutations in FRI lead to early flowering due to reduced FLC expression, but other genetic pathways (e.g., vernalization, autonomous pathways) can also modulate FLC levels independently of FRI (Jones et al., 2024). In addition, we cannot exclude the possibility that another gene, possibly multiple BnaFRI-like genes or another previously unidentified homologue of BnaFRI, could compensate for the loss of function at BnaFRI in these lines. As BnaFLC is not the target of BnaFRI protein function, a comparative transcriptome analysis of B. napus with and without BnaFRI protein may reveal which genes, if any, are transcriptionally regulated by BnaFRI.
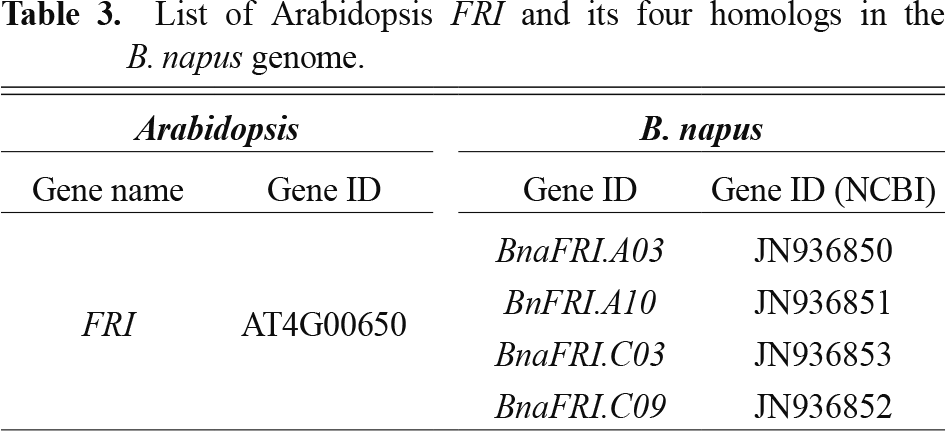

List of Arabidopsis FRI and its four homologs in the B. napus genome.
Heterologous expression of an Arabidopsis FLC transgene in an early flowering B. napus line resulted in a substantially delayed floral transition (Tadege et al., 2001). Interestingly, expression of AtFLC in B. napus affected flower development and fertility, suggesting possible functional differences between A. thaliana FLC and B. napus FLC (BnFLC) homologs. In addition, transformation of five individual BnFLC homologs, BnFLC1~BnFLC5 commonly resulted in delayed floral transition when compared to those of an Arabidospis wild type accession, Ler (Tadege et al., 2001). Genome wide association studies (GWAS) analysis on flowering time variation using 182 B. napus accessions collected from different geological locations around the world revealed that several SNPs (single nucleotide polymorphisms) associated with flowering time regulation were highly linked to candidate genes like FT, FRUITFUL (FUL), FLC, CO, FRI, PHYTOCHROME B (PHYB) homologs within the B. napus genome (Raman et al., 2016). The genome of allopolyploid Brassica napus (rapeseed) seems highly duplicated and it contains nine annotated copies of Bna.FLC (Schiessl et al., 2019) (Table 2). Only three Bna.FLC copies including Bna.FLC.A03b, Bna.FLC.A10, and to some degree Bna.FLC.C02, were differentially expressed between the winter type and spring type B. napus lines. In addition, expression levels of the other six Bna.FLC copies were not correlated with the vernalization requirement. Three of the six Bna.FLC copies including Bna.FLC.C03a, Bna.FLC.C09a, Bna.FLC.C09b were not responsive to vernalization treatment. Meanwhile, Bna.FLC.C03b was not expressed at all, indicating it is a pseudogene.
Interestingly, several independent studies commonly reported that transposable element (TE) insertion in the genomic region of Bna.FLC resulted in different flowering behavior upon vernalization (Akter et al., 2021). For instance, a 621-bp insertion of TE within the upstream region of BnaFLC.A10 was associated with impairment of vernalization-mediated suppression of BnaFLC.A10 in B. napus (Hou et al., 2021). This TE insertion was commonly detected in 73% of winter type B. napus lines (449 out of 619 lines) (Yin et al., 2020). Another 4,422-bp TE insertion was detected in 57% of semi-winter type B. napus lines, resulting in the malfunction of Bna.FLC.A10 and earlier flowering than winter type B. napus. Furthermore, a 5,625-bp TE insertion in the first exon of BnaFLC.A10, an 810-bp TE insertion in the 7th exon of BnaFLC.A02, a 2,833-bp TE insertion in the first intron of BnaFLC.A02 were found in spring type B. napus, suggesting a close connection between TE insertions on the BnaFLC and vernalization responsiveness in B. napus plants (Chen et al., 2018a; Yin et al., 2020). However, another study proposed that overall BnaFLC transcript dynamics, rather than the transcript level of specific BnaFLC, best explain flowering behavior in response to vernalization in B. napus cultivars (Calderwood et al., 2021).
Using linkage and association studies, VIN3 was shown to contribute to flowering-time variation (Han et al., 2022; Shah et al., 2018). The B. napus genome contains four B. napus VIN3 homologs (BnaVIN3.A02, BnaVIN3.A03, BnaVIN3.C02, and BnaVIN3.C03). Expression levels of all BnaVIN3 homologs were induced by cold in both leaf and apex, similar to Arabidopsis, and were substantially downregulated when plants were return to warm growth temperatures (Schiessl et al., 2019). Another study showed that exposure to transient warm temperatures significantly reduced cold-induced transcript levels of these B. napus VIN3 homologs in B. napus (O’Neill et al., 2019), indicating that a cold temperature signal affects expression of BnaVIN3 homologs, as expected from Arabidopsis VIN3. Among them, BnaVIN3.C03 exhibited the lowest transcript level, while other BnaVIN3 homologs displayed comparable transcript levels. A study was done using genome-wide marker distribution analysis on 35 flowering regulators in 280 diverse B. napus genotypes encompassing four different eco-geographical morphotypes (winter, semi-winter, spring, and swede types). The study found that BnaVIN3.A02 is a major candidate for causing the winter-spring split, and BnaVIN3.A03 for the swede split (Schiessl et al., 2017). However, no non-synonymous SNP (single nucleotide polymorphism) was found to be correlated this categorization.
VIN3 is induced by vernalization and plays a role as a master regulator for floral transition in Arabidopsis (Sung and Amasino, 2004a). Similarly, it is likely that BnVIN3 homologs play a role as transcriptional repressors in cooperation with PRC2 and PRC1 complexes for epigenetic suppression of BnFLC homologs. During long-term cold, BnFLC homologs are epigenetically suppressed by BnVIN3-containing PRC2/1 complexes, and this event eventually triggers de-repression of downstream floral integrator genes like BnFT and BnSOC1 homologs, which promote the floral transition of B. napus (Fig. 1C).
It could be interesting to explore whether BnaVIN3s mediate chromatin modifications on Bna.FLC in response to vernalization. In addition to the vernalization response, it was previously reported that Arabidopsis VIN3 is significantly upregulated in a hypoxia condition (Bond et al., 2009). To our knowledge, it has not been clarified whether environmental signals like hypoxia and temperature fluctuation differently affect expression of BnaVIN3 homologs. For in-depth investigations into the functional role of different BnaVIN3 homologs in B. napus plants, development of knock-out mutants of individual BnaVIN3 homologs may be required. This may be possible utilizing a CRISPR/Cas9 tool to modify BnaVIN3 expression, which could lead to the modulation of flowering behavior of B. napus plants.
Brassica junceaBrassica juncea (Indian mustard) is derived from the interspecific crossing of Brassica rapa (AA, 2n = 20) and Brassica nigra (BB, 2n = 16), so it is an allotetraploid (AABB, 2n = 36) (Table 2). Genomic polyploidization resulted in complex, highly repetitive flowering regulatory genes, thus making it difficult to elucidate the molecular mechanism underlying the floral transition in B. juncea. Because flowering behavior directly affects crop productivity and economic benefits, understanding flowering regulatory networks and characterization of major flowering time regulator genes in B. juncea is important to manipulate flowering time.
Flowering of B. juncea has been shown to be affected by both photoperiod and vernalization. Studies have reported that longer days (long-day photoperiod) promotes floral transition, while short days delay flowering. This suggests that flowering time of B. juncea plants is influenced by the duration of light exposure (Kang et al., 2021; Macioszek et al., 2021). In addition, a study showed that East European germplasms are adapted to long-day conditions, while many Chinese winter-type mustards are naturally biennial, requiring vernalization for floral transition. In contrast, Indian germplasms are winter annuals with a very short vegetative phase and typically do not require vernalization (Akhatar et al., 2021). Thus, it is likely that both photoperiod and vernalization can affect the floral transition of B. juncea, even though there may be differences in the degree of response.
A genome-wide computational analysis of the flowering-related gene identified 179 genes that were flowering regulatory genes in B. juncea. Among them, 84 genes were mapped to the B. juncea A sub-genome, while the other 79 genes were mapped onto the B. juncea B sub-genome (Yang et al., 2014b). Expression pattern analysis of these flowering-related genes suggested that BjuACO4, BjuAFT1, BjuBFT4, BjuASOC1, BjuASOC4 may play a key role in regulating the floral transition in B. juncea. In addition, the B. juncea genome was shown to contain five CO homologs (BjuACO1, BjuACO2, BjuBCO3, BjuACO4, and BjuBCO5). A functional study using BjuACO4 and BjuBCO3-silenced transgenic lines provided new evidence for the genetic interactions between BjuCOs and BjuFLCs (Zhao et al., 2019). Thus, it is likely that expression of floral repressors, that is, BjuFLCs, may be modulated not only by vernalization pathway regulators, but also photoperiod flowering pathway regulators, because BjuFLCs expression levels were significantly down-regulated under long-day treatments. This is in a line with a previous report indicating that floral transition of B. juncea can be influenced by both a BjuCOs-mediated photoperiod pathway, as well as a vernalization pathway (Fig. 1D). Taken together, genomic polyploidization resulted in a much more complicated flowering regulatory network in B. juncea plants.
Flowering time is an agriculturally important trait. Knowledge of the mechanisms underlying flowering time control in plants can be applied to improve important crop traits. For example, the leafy vegetables of the genus Brassica, such as Chinese cabbage, eventually lose their commercial value after bolting because energy and metabolites are reallocated to reproductive tissues, causing a reduction in their leafy tissues. In contrast, canola (B. napus) is mainly cultured to harvest seeds for oil. Therefore, appropriate control of flowering time can maximize the productivity and quality of leaves, flowers, or seed tissues in crop plants. In this regard, understanding the molecular mechanisms underlying floral transition is of particular interest in agricultural breeding programs.
This review described the current understanding of the molecular mechanisms revealed in the model plant Arabidopsis and several crop plants. Intensive studies on this subjecy have treatly increased our knowledge of the genes involved in these pathways, protein-protein interactions, and the molecular interaction networks among these genes. In addition, the epigenetic chromatin regulators (i.e., PRC complexes) of the key genes involved in flowering time have expanded our understanding of how plants optimize their growth and development according to changing environmental cues. However, some areas remain unclear. For example, it is not fully understood how plants accurately measure the duration of cold in the winter season and prevent premature flowering even with temperature variations during winter. The mechanisms by which plants can assess the length of vernalizing cold temperatures is likely to be independent of the cold acclimation pathway.
Even though our understanding of flowering pathways in crop plants has been greatly enhanced, the molecular mechanisms underlying these pathways remain poorly understood. Therefore, further research is required to explore and elucidate the molecular mechanisms controlling flowering pathways in crop plants. We expect that comparative studies based on the model plant Arabidopsis will help us acquire a more comprehensive understanding of the molecular details underlying plant flowering programs. Especially, computation analysis using next-generation sequencing data and genomics tools for map-based cloning will highly accelerate the identification of agriculturally important loci and genes in many crop species. As a result, these current approaches will help us identify the essential DNA elements required for the vernalization-mediated floral transition. Furthermore, a recently developed genome-editing tool, CRISPR-Cas9, can be utilized to modify the identified DNA elements to engineer crop traits, including flowering time, to enhance the commercial value of crop plants.
I apologize to colleagues whose work could not be included owing to space constraints.