2017 Volume 93 Issue 7 Pages 498-510
2017 Volume 93 Issue 7 Pages 498-510
Estrogen receptors (ER) are important transcription factors to relay signals from estrogen and to regulate proliferation of some of breast cancers. The cycling of estrogen-induced DNA binding and ubiquitin-linked proteolysis of ER potentiates ER-mediated transcription. Indeed, several transcriptional coactivators for ER-dependent transcription ubiquitinate ER. Histone acetyltransferase (HAT) Hbo1/KAT7/MYST2, involved in global histone acetylation, DNA replication, transcription, and cellular proliferation, promotes proteasome-dependent degradation of ERα through ubiquitination. However, molecular mechanism for ubiquitination of ERα by Hbo1 is unknown. Here we report the intrinsic ubiquitin E3 ligase activity of Hbo1 toward the ERα. The ligand, estradiol-17β, inhibited E3 ligase activity of Hbo1 for ERα in vitro, whereas hyperactive ERα mutants from metastatic breast cancers resistant to hormonal therapy, were better substrates for ERα ubiquitination by Hbo1. Hbo1 knock-down caused increase in ERα expression. Hbo1 is another ERα coactivator that ubiquitinates ERα.
Steroid hormone estrogen has numerous physiological functions, including reproduction, bone homeostasis, cardiovascular health, behavior, cognition, and cellular proliferation. Estrogen transmits its signal into the cell through association with estrogen receptors (ERα and ERβ). The ligand-ERα complex directly or indirectly binds to DNA, recruits co-regulatory protein complexes, and activates or inactivates transcription.1)–4) In addition to genomic action, estrogen exerts its function through binding to DNA-binding transcription factors or through membrane-bound ERα. Moreover, ERα action is modulated by co-regulators,5),6) splicing,7) miRNAs,8) long non-coding RNAs,9) and posttranslational modifications.10) Clinically important, two-thirds of breast cancers are positive for ERα and respond to hormonal therapy.11)
The ubiquitin-proteasome pathway12) is responsible for basal and ligand-induced ERα degradation.13)–15) Counterintuitively, this ligand-induced ERα degradation, potentiates transcriptional activation by ERα through cycling of DNA binding and ubiquitin-dependent proteolysis of ERα. ERα co-activators that ubiquitinate E3 include E6-AP, Mouse double minute 2 homologue (Mdm2), the breast cancer susceptibility gene 1/Brca1-associated RING domain protein (BRCA1/BARD1) estrogen-responsive finger protein (EFP), and SKP1–CUL1–F-box protein complex (SCF). In addition, growth-related signal pathways, such as mitogen-activated kinase (MAPK) and phosphoinositide 3-kinase (PI3K) pathways, stimulate ubiquitination of ERα through ERα phosphorylation.16)
Hbo1, a histone acetyltransferase binding to origin recognition complex (ORC1),17) regulates histone acetylation,18)–20) replicational licensing,21),22) origin activation,23) adipogenesis,24) pluripotency and self-renewal of embryonic stem cells,25) and chromatin assembly of centromeres.26) Hbo1 is linked to cellular proliferation18),27) and subject to functional modulation upon DNA damage.28),29) Hbo1 belongs to MYST family of histone acetyltransferases.30) We have found that Hbo1 promotes ubiquitin-mediated degradation of ERα and modulates ERα-mediated transcription and proliferation.31) Hbo1 appears to be a coactivator that ubiquitinates ERα. However, it is unclear whether acetylation of ERα by Hbo1 facilitates ERα ubiquitination or Hbo1 directly ubiquitinates ERα.
Here we investigated molecular mechanism of the promoted ubiquitination of ERα by Hbo1. The ligand-binding domain (LBD) of ERα was ubiquitinated by Hbo1. Hbo1 is an atypical ubiquitin E3 ligase for ERα. The self-ubiquitination activity of Hbo1 was mapped to the MYST domain. The ligand, estradiol-17β, inhibited E3 ligase activity of Hbo1 toward ERα. In contrast, hyperactive ERα mutants found in metastatic breast cancers resistant to hormonal therapy, were better substrates for ubiquitination by Hbo1. Irrespective of the presence and absence of the ligand, knocking down Hbo1 expression caused increase in ERα expression. These results suggest that Hbo1 is a novel E3 ubiquitin ligase that activates ERα-dependent transcription.
17β-estradiol was purchased from Sigma-Aldrich (St. Louis, MO, USA), MG-132 from Enzo Life Sciences (Farmingdale, NY, USA), Cy5-conjugated Streptavidin from Jackson ImmunoResearch Laboratories (West Grove, PA, USA), and Coomassie Stain Solution from Bio-Rad (Hercules, CA, USA). The recombinant KAT7 (Hbo1 purified from Sf9 cells), MurF1, GST-tagged p53, and ubiquitin proteins were purchased from Active Motif (Carlsbad, CA, USA), R&D systems (Minneapolis, MN, USA), EMD Millipore (Temecula, CA, USA) and UBPBio (Aurora, CO, USA), respectively. Anti-Hbo1 antibody was described.17) The following commercial antibodies were purchased: anti-FLAG monoclonal and anti-α-tubulin monoclonal from Sigma-Aldrich (St. Louis, MO, USA), anti-HA monoclonal from Roche (Basel, Switzerland), anti-ERα polyclonal from Santa Cruz (Dallas, TX, USA), anti-ubiquitin polyclonal from Cell Signaling Technology (Danvers, MA, USA), and anti-GST polyclonal and anti-His-tag monoclonal from Medical and Biological Laboratories (Nagoya, Japan).
Mutations in the ERα gene and purification of recombinant proteins.Deletion mutants of the human ERα gene31) were made by amplifying corresponding DNA fragments using PCR. Base substitutions were made using PrimeStar Mutagenesis Basal kit (TaKaRa Biotechnology, Shiga, Japan) and the mutations were verified by sequencing. Recombinant proteins were expressed in bacteria BL21(DE3)pLysS. GST-tagged ERα LBD (amino acids: 304–554) and GST-tagged human Mdm2 (a gift from Dr. Koji Okamoto, National Cancer Center Research Institute, Tokyo, Japan) were purified with Glutathione Sepharose.32) Throughout the GST-ERα LBD purification, estradiol-17β was not included in the growth media or buffers for stabilization of ERα LBD,33) to assay the effect of ligand during the ubiquitination reaction. The His-tagged full-length human Hbo1 (amino acids: 1–611) protein was purified on Heparin Sepharose (GE Healthcare, Little Chalfont, UK), whereas His-Hbo1 (amino acids: 225–611) and His-Hbo1 (amino acids: 311–611) proteins28) were purified on Ni-NTA Agarose (Qiagen, Hilden, Germany).
Ubiquitination assay.In vivo ubiquitination assay was performed as previously.31) For in vitro ubiquitination assay, reactions were carried out using a ubiquitinylation kit purchased from Enzo Life Sciences (Farmingdale, NY, USA) containing 2.5 µM biotinylated or non-tagged ubiquitin, 100 nM His-tagged E1, approximately 1–2.5 µM His-tagged E2, and 5 mM Mg-ATP, and 20 U/mL yeast inorganic pyrophosphatase (New England Biolabs, Ipswich, Massachusetts, USA). Detection was made by immunoblotting with Cy5-conjugated streptavidin (Cell Signaling Technology) or anti-ubiquitin antibody. His-tagged UbcH5c was used for Figs. 2A and 2E, whereas non-tagged UbcH5c (R&D Systems, Minneapolis, MN, USA) was used for all the other ubiquitination reactions. To affinity-purify His-tagged and GST-tagged proteins, stringent washing conditions were completed using 0.3 M NaCl and 0.1% Triton-X for Figs. 3D and 3E, and 0.5 M NaCl plus 0.1% Triton-X for Figs. 4, 5A, and 5D, respectively.
siRNA knock-down and quantitative reverse transcription (qRT)-PCR.siRNA and qRT-PCR experiments were as described previously.34) siRNAs for Hbo1 targeted nucleotide 92 of the coding sequences of Hbo1.31) For qRT-PCR analysis, data was normalized to glyceraldehyde-3-phosphate dehydrogenase (GAPDH) expression. Values were expressed as mean ± SD (n = 4, unless otherwise stated). The PCR primer sets for Hbo1 and GAPDH were described.31)
To map sites of ERα ubiquitination promoted by Hbo1, deletion mutants of ERα were constructed (Fig. 1). FLAG-tagged ERα mutants, Hbo1, and HA-ubiquitin were transfected into 293T cells, followed by immunoprecipitation with anti-FLAG antibody under denaturing conditions, and immunoblotted with anti-HA and anti-FLAG antibodies (Fig. 1B). High-molecular-weight ubiquitinated proteins were detected in mutants 1, 2, and 3 (lanes 3–5 and 8–10), comparable to the full-length ERα (lanes 2 and 7), suggestive of extensive ubiquitination in ERα. The ubiquitinated signals of mutants 2 and 3 were stronger than that of mutant 1 (lanes 4 and 5 compared to lane 3; lanes 9 and 10 compared to lane 8). This result indicated that the hinge region (H) and the ligand-binding domain (LBD), shared by mutants 2 and 3 of ERα, were preferentially ubiquitinated by Hbo1. As lysines 302 and 303 in the hinge region protect unliganded ERα turnover,35) we predicted that the removal of the hinge region would boost ubiquitination of ERα. Deletion mutant 4 (amino acids: 304–554) was constructed, mostly composed of LBD, and assayed for ubiquitination by Hbo1 (Fig. 1C). Mutant 4 gave a stronger poly-ubiquitination signal than the full-length ERα (lanes 3 and 6). These results suggested that Hbo1 promotes a robust ubiquitination of the ERα LBD. From this point forward, we focused on the analysis of ERα LBD.

Ubiquitination of ERα LBD stimulated by Hbo1. (A) Domain structure of the ERα protein. DBD: DNA-binding domain; LBD: ligand-binding domain; H: hinge region. Full and mutants 1 through 4, cover amino acids 1–595, 1–298, 282–595, 1–179 plus 269–595, and 304–554 of the ERα protein, respectively. Expected molecular weights are shown. (B–C) FLAG-tagged ERα mutants, Hbo1, and HA-ubiquitin were transfected into 293T cells. The cells were treated with MG-132 for 6 hours, lysed under denaturing conditions, and immunoprecipitated with anti-FLAG antibody. The immunoprecipitates were blotted with anti-HA or anti-FLAG antibodies. v: empty FLAG vector; Full: FLAG-tagged full-length ERα.
To test whether Hbo1 alone has ubiquitin E3 ligase activity, an array of E2 ubiquitin conjugating enzymes for self-ubiquitination of commercial Hbo1 protein were screened by mixing commercial recombinant Hbo1 protein with E1, E2, ATP, and biotinylated ubiquitin and probing with Cy5-conjugated streptavidin (Fig. 2A). High-molecular-weight ubiquitination bands greater than 250 kDa (Ubn-E3) were detected by using UbcH5b and UbcH5c as an E2 (lanes 12 and lane 14), similar to the control MurF1, a known E3 ligase36) (lane 2), suggesting self-ubiquitination of Hbo1. The lack of signal by immunoblotting of Hbo1 protein with anti-ubiquitin antibody (Fig. 2B, lanes 4 and 5) ruled out a possibility that the original preparation of Hbo1 protein was ubiquitinated. To confirm the E2 screen result by a different assay, we incubated the recombinant Hbo1 protein with E1, E2 (UbcH5c), ATP, and non-tagged ubiquitin and immunoblotted the reaction mixture with anti-ubiquitin antibody (Fig. 2C). High molecular-weight poly-ubiquitination signals were detected when Hbo1 was used as an E3 enzyme (lane 3), comparable to the positive control, MurF1 (lane 2), indicating a self-ubiquitination activity of Hbo1. Equal loading of E1 was confirmed by blotting with anti-His antibody (bottom panel, Fig. 2C). When any one component of the ubiquitin reaction was omitted (Fig. 2D), high molecular weight poly-ubiquitination was abolished. Hbo1 signal was diminished only when all the necessary components were included (bottom panel, lane 5). These results demonstrated dependency of the reaction on E1, E2, and ubiquitin. Similar experiment was done by incubation of biotinylated ubiquitin, His-tagged E1, His-tagged E2 (UbcH5c), and commercial Hbo1, followed by removal of E1 and E2 by mixing with Ni-NTA Agarose. The supernatant was probed for ubiquitin by immunoblotting with Cy5-streptavidin (Fig. 2E). High-molecular-weight ubiquitinated signals were detected only when all four components were included (lane 2). All these results suggested E3 ligase activity of Hbo1.

Self-ubiquitination of Hbo1. (A) E2 screen for Hbo1 E3 ligase assay. Ten different His-tagged E2s were incubated with (+) or without (−) commercial recombinant Hbo1 in the presence of His-tagged E1, ATP, and biotinylated ubiquitin. The reaction mixture was run on SDS-PAGE, transferred onto the membrane, and probed with Cy5-conjugated streptavidin to detect ubiquitin. Recombinant MurF1 protein (1.25 µg) served as a positive control for E3 ligase activity (lanes 1 and 2). (B) Recombinant Hbo1 protein from insect cells (lanes 1 and 4) and from bacteria (lanes 2 and 5), and recombinant ubiquitin (lanes 3 and 6), were blotted with anti-Hbo1 or anti-ubiquitin antibodies. Ub: ubiquitin. (C) Commercial recombinant Hbo1 protein was incubated with His-tagged E1, E2 (UbcH5c), ATP, and non-tagged ubiquitin. The reaction mixture was run on SDS-PAGE, transferred onto the membrane, and probed with anti-ubiquitin or anti-His antibodies. Recombinant MurF1 protein (1.25 µg) served as a positive control (lane 2). Ub: ubiquitin. (D) Ubiquitination reaction mixture performed in the listed combination using commercial Hbo1 protein and non-tagged ubiquitin, was blotted with anti-ubiquitin or anti-Hbo1 antibodies. (E) Hbo1 protein from Sf9 cells (1.2 µg) was incubated in the listed combinations for 3 hours. After removal of His-tagged E1 and E2 by mixing with Ni-NTA Agarose, the reaction mixture was blotted with Cy5-conjugated streptavidin to detect ubiquitin.
Although Hbo1 lacks RING domains, HECT domains, and RBR domains, which are characteristics of E3 ligases,37) it possesses two zinc-finger like motifs: one within the MYST domain and the other outside the MYST domain (Fig. 3A) like other atypical ubiquitin E3 ligases.38)–40) To map the E3 ligase activity of Hbo1, deletion mutants of Hbo1 were bacterially purified (Fig. 3B). No signal by immunoblotting with anti-ubiquitin antibody against the purified Hbo1 deletion mutants (lanes 2 and 3, Fig. 3C) demonstrated lack of ubiquitin modification on those Hbo1 deletion mutants. We incubated His-tagged Hbo1 mutants with His-tagged E1, non-tagged E2, biotin-ubiquitin, and ATP, followed by affinity purification of His-tagged Hbo1 and E1 proteins, with Ni-NTA Agarose under stringent condition. The purified proteins, were probed with Cy5-streptavidin and anti-His antibody (Figs. 3D and 3E). Immunoblotting with anti-His antibody detected E1 in all lanes except for lane 2, where no E1 was included (bottom panels, Figs. 3D and 3E). On the other hand, Cy5-streptavidin blot revealed that the ubiquitinated E1 (approximately 120 kDa band) was detected in lanes 1, 3, and 5 (Ub-E1), where E1 and ubiquitin were included (top panels, Figs. 3D and 3E). As expected, the ubiquitination of E1 was dependent on ubiquitin and E1. In Fig. 3D, in Cy5-strepavidin blotting, bands were detected for His-Hbo1 (225–611), predominantly around 80 kDa in size, migrating slower than the non-modified His-tagged Hbo1 (225–611) solely in lane 5 where all the necessary components were included (bracketed). Those bands reacted with anti-His antibody (lane 10, arrowhead), suggesting that Hbo1 (225–611) has E3 ligase activity. In contrast, Hbo1 (311–611) produced an approximately 40 kDa band when all reagents were included (lanes 5 and 10, Fig. 3E), detected by both ubiquitin and His blots (arrowheads, Fig. 3E), suggestive of mono-ubiquitination. These results suggest that the MYST domain of Hbo1 has E3 ligase activity. The ubiquitin E3 ligase activity detected in bacterially purified Hbo1 protein eliminated a possibility that contaminants in commercial Hbo1 from Sf9 cells had ubiquitin E3 activity.

MYST domain of Hbo1 has ubiquitin E3 ligase activity. (A) Schematic representation of Hbo1 domains and deletion mutants. Z: zinc finger; MYST: MYST domain. Expected molecular weights are shown. (B) Coomassie staining of His-tagged Hbo1 (amino acids: 225–611) and His-Hbo1 (amino acids: 311–611) proteins. (C) His-tagged E1, His-Hbo1 (225–611), His-Hbo1 (311–611), and non-tagged ubiquitin proteins were immunoblotted with anti-ubiquitin or anti-His antibodies. (D, E) Following ubiquitination reaction in the listed combination, His-tagged E1 and Hbo1 proteins (Hbo1 (225–611) for (D) and Hbo1 (311–611) for (E)) were affinity-purified with Ni-NTA Agarose under stringent condition, run on SDS-PAGE, and blotted with Cy5-conjugated streptavidin or anti-His antibody. Arrowheads: ubiquitinated Hbo1 deletion mutants.
Prior to examining whether Hbo1 ubiquitinates ERα LBD, to confirm that our assay system for ubiquitination is appropriate, we checked whether Mdm2 ubiquitinates p53 in vitro41) by our assay. We incubated His-E1, E2 (UbcH5c), GST-Mdm2 protein, biotinylated ubiquitin, ATP, and GST-tagged p53, followed by affinity-purification of GST-tagged proteins under stringent condition. The GST-tagged Mdm2 and p53 proteins were analyzed by immunoblotting with Cy5-streptavidin and anti-GST antibody (Fig. 4A). No poly-ubiquitination of GST-p53 was observed without Mdm2 (lanes 1 and 4). On the other hand, high-molecular-weight poly-ubiquitination signal (lanes 3 and 6) confirmed that Mdm2 has intrinsic E3 ligase activity. When GST-p53 was incubated with GST-Mdm2, in addition to high-molecular-weight poly-ubiquitination signal, smeared bands ranging between 75 kDa and 100 kDa (bracketed) was detected (lanes 2 and 5), suggestive of ubiquitinated GST-p53. Thus, our assay could detect ubiquitinated substrates.

Ubiquitination of ERα LBD by Hbo1. (A) GST-tagged p53 (2 µg) and GST-tagged Mdm2 (E3 ligase) were mixed in the indicated combination in the presence of E1, E2 (UbcH5c), biotinylated ubiquitin, and ATP for 2 hours, followed by affinity purification of GST-tagged p53 and Mdm2 proteins with Glutathione Sepharose under stringent condition and by immunoblotting with Cy5-strepavidion (left panel) or anti-GST antibody (right panel). Arrowhead: ubiquitinated E1. Bracket: ubiquitinated GST-p53. (B) Commercial Hbo1 protein (1.2 µg) was incubated in the listed combinations. GST-tagged ERα LBD protein was affinity-purified with Glutathione Sepharose under stringent condition, and assayed for ubiquitin (left panel) and GST (right panel). (C) No enzyme (lane 1), Recombinant His-Hbo1 (amino acids: 311–611) protein (lane 2: 1 µg; lane 3: 2.5 µg), and commercial Hbo1 (1.2 µg, lane 4) were assayed in the ubiquitination reaction, and GST-ERα LBD protein was stringently affinity-purified with Glutathione Sepharose. The purified proteins were separated, transferred onto membrane, and ubiquitin and GST were detected with Cy5-conjugated streptavidin (left panel) or with anti-GST antibody (right panel), respectively. The identity of the band between 50 and 75 kDa ($\ast$) is unknown.
A test was performed to determine whether Hbo1 ubiquitinates ERα LBD in vitro. We incubated His-E1, E2 (UbcH5c), the commercial Hbo1 protein, biotinylated ubiquitin, ATP, and GST-tagged ERα LBD, followed by affinity-purification of GST-tagged ERα LBD protein under stringent condition. The GST-ERα LBD proteins were analyzed by immunoblotting with Cy5-streptavidin and anti-GST antibody (Fig. 4B). High-molecular-weight ubiquitinated proteins were only observed when all the necessary components were included (lanes 6 and 12). This result indicated that Hbo1 per se could ubiquitinate ERα in vitro. Next the mutant Hbo1 (311–611) was examined to determine whether it could ubiquitinate ERα LBD (Fig. 4C). Use of Hbo1 (311–611) protein as an E3 enzyme lacked a high-molecular-weight ubiquitination (lanes 2, 3, 6, and 7), suggesting that the MYST domain was not sufficient for ERα LBD ubiquitination by Hbo1. These results indicated that the N-terminal portion of Hbo1 was required for E3 ligase activity toward ERα LBD.
Estradiol-17β inhibited ERα ubiquitination by Hbo1 in vitro.The ligand destabilizes ERα protein in a proteasome-dependent manner.14),15),42) The association of ERα LBD with the ligand, estradiol-17β, involves multiple amino acids via hydrogen bonds and hydrophobic contacts.43) To examine the effect of estradiol-17β on ERα ubiquitination by Hbo1, estradiol-17β was included in the ubiquitination reaction, followed by affinity-purification of GST-ERα LBD, and assayed for ubiquitin by immunoblotting (Fig. 5A). To negate an effect of ligand carryover from ERα LBD, estradiol-17β was not used to stabilize ERα LBD throughout the purification procedures of GST-ERα LBD. With the increase of estradiol-17β, high-molecular-weight ubiquitination signal of ERα was reduced in a dose-dependent manner (lanes 3 and 4 compared to lane 2), suggesting that estradiol-17β inhibited ERα ubiquitination by Hbo1 in vitro. As the ERα LBD bound tightly to ubiquitin,44) ubiquitinated E1 (about 120 kDa) was pulled down concomitantly with GST-ERα LBD, even under stringent conditions (arrowhead).
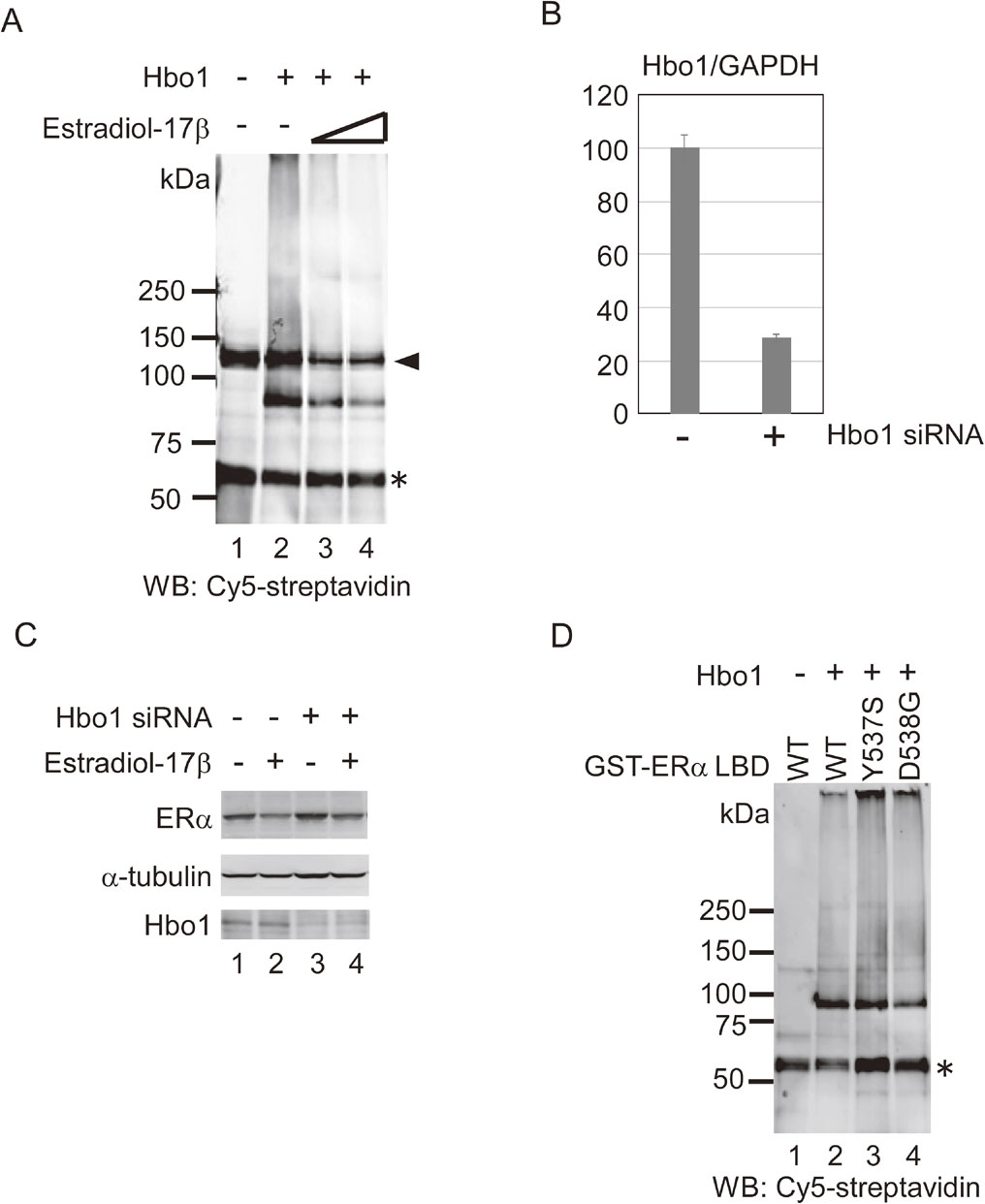
Modulation of Hbo1 E3 ligase activity. (A) Commercial Hbo1 protein (1.2 µg) was incubated with GST-ERα LBD in the ubiquitination reaction in the presence of estradiol-17β (lane 2: vehicle; lane 3: 20 nM; lane 4: 200 nM). GST-tagged ERα LBD protein was purified and probed with Cy5-conjugated streptavidin to detect ubiquitin. The identity of the band between 50 and 75 kDa ($\ast$) is unknown. Arrowhead: ubiquitinated E1. (B) Expression of Hbo1 mRNA was analyzed by qRT-PCR following Hbo1 knock-down. A ratio of the Hbo1 mRNA to GAPDH mRNA in the control was arbitrarily set to 100. Each value was represented as a mean ± SD. (C) MCF-7 cells were transfected with Hbo1 siRNA for 48 hours. Post-transfection, the cells were treated with or without 10−8 M estradiol-17β for 24 hours. The cell lysates were probed with anti-ERα, anti-α-tubulin and anti-Hbo1 antibodies. (D) Commercial Hbo1 protein (1.2 µg) was incubated with GST-ERα LBD (lane 2), GST-ERα LBD-Y537S (lane 3), and GST-ERα LBD-D538G (lane 4) in the ubiquitination reaction. GST-tagged ERα LBD protein were analyzed as in Fig. 5A. Asterisk ($\ast$): origin unknown.
Based on ligand-inducible degradation of ERα,14),15),42) degradation of ERα by Hbo1,31) and inhibition of E3 ligase activity of Hbo1 for ERα by the ligand (Fig. 5A), we examined whether Hbo1 was involved in basal or estrogen-induced turnover of ERα. We knocked down expression of Hbo1 in a breast cancer cell line, MCF-7 cells, cultured the cells with or without estradiol-17β, and examined expression of Hbo1 by qRT-PCR analysis (Fig. 5B) and of ERα by western blotting (Fig. 5C). A 72% reduction of Hbo1 mRNA expression by siRNA knock-down was achieved (Fig. 5B). In accord with the qRT-PCR result, Hbo1 protein decreased upon siRNA knock-down of Hbo1 (bottom panel, Fig. 5C). An equivalent loading of proteins onto each lane was judged by probing with anti-α-tubulin antibody (second panel, Fig. 5C). ERα expression (upper panel, Fig. 5C) was decreased upon estrogen treatment (lane 1 compared to lane 2) due to ligand-mediated turnover of ERα as previously reported.14),15) With no ligand, depletion of Hbo1 caused an increase in ERα protein (lane 1 compared to 3), consistent with the previous study.31) When treated with the ligand, ERα expression increased in Hbo1 siRNA cells compared to that of the siRNA control (lanes 2 and 4). These results suggest that Hbo1 regulates the stability of ERα, irrespective of the presence and absence of the ligand.
Enhanced ubiquitination of hyperactive ERα mutants by Hbo1.ERα somatic mutations in LBD, ERα-Y537S and ERα-D538G, found in ER-positive metastatic breast cancers, activate transcription even in the absence of ligand, providing a mechanism for resistance to hormonal therapy.45)–49) Due to the linkage of ERα ubiquitination to transcriptional activation,16) constitutive transcriptional activation by the hyperactive ERα mutants might be enhanced by ERα ubiquitination. To determine whether these hyperactive ERα mutants were good substrates of in vitro ubiquitination by Hbo1, we introduced the mutations by site-directed mutagenesis. The recombinant GST-tagged ERα LBD proteins were examined for ubiquitination by Hbo1 (Fig. 5D). The ERα LBD-Y537S and ERα LBD-D538G proteins were ubiquitinated by Hbo1 (lanes 3 and 4) more than the wild-type (lane 2). Notably, the ERα LBD-Y537S showed a more robust ubiquitination than the wild-type, whereas ERα LBD-D538G showed a modest increase in ubiquitination.
Here, we report analysis of ERα ubiquitination by Hbo1. ERα LBD is preferentially ubiquitinated by Hbo1. Hbo1 has intrinsic ubiquitin E3 ligase activity for ERα LBD. We discovered two factors which regulates E3 ligase activity of Hbo1 for ERα: the ligand, estradiol-17β and the hyperactive ERα mutants derived from metastatic breast cancers. The ligand inhibited E3 ligase activity of Hbo1 for ERα, whereas the hyperactive ERα mutants were better substrates for the E3 ligase activity. Hbo1 regulated basal and estrogen-induced turnover of ERα.
Ubiquitination of ERα hyperactive mutants by Hbo1.The hyperactive ERα mutants, ERα LBD-Y537S and ERα LBD-D538G, which activate ERα-dependent transcription in the absence of the ligand, were ubiquitinated more than the wild-type ERα by Hbo1 (Fig. 5D). Of interest, the marked difference in ubiquitination between the ERα LBD-Y537S and ERα LBD-D538G mutants was similar to that of transcriptional activation by ERα-Y537S and ERα-D538G in the absence of the ligand (Fig. 345) and Fig. 2a46)), respectively, suggesting that ERα ubiquitination by Hbo1 contributed to transcriptional activation by ERα. It is speculated that the enhanced ubiquitination of the ERα mutants by Hbo1 induces cyclic proteasome-mediated turnover of ERα, leading to potentiation of estrogen signaling.50) Ubiquitin-mediated degradation of ERα by Hbo1, in cooperation with histone acetylation by Hbo1, may lead to transcriptional activation. ERα-dependent genes, such as E2F1, RRM2, CTSD, and NRGM, which are activated by Hbo1,31) may be subjected to dual enhancement by its two enzymatic activities, HAT and ubiquitin E3 ligase. In this context, Hbo1 might contribute to enhancing hormonal resistance of breast cancers with ERα mutations.
Inhibition of Hbo1-mediated ERα ubiquitination of by the ligand.It seems puzzling that Hbo1 promoted ligand-induced turnover of ERα, although the ligand, estradiol-17β, inhibits Hbo1 E3 ligase activity in vitro. One explanation is that the interaction of the ligand with the ERα LBD masks LBD, facilitates ubiquitination at regions other than LBD by Hbo1, and induces subsequent proteasome-dependent degradation. Thus, the binding of the ligand could enhance ERα ubiquitination by Hbo1. The extensive ubiquitination of ERα by Hbo1 (Fig. 1B) supports the notion. Lysines 302 and 303 in the hinge region are important for ligand-induced degradation of ERα35) and may be target sites for ubiquitination by Hbo1.
Possible E3 ligase activity of MYST HATs.Several HATs have ubiquitination-linked enzymatic activity. p300/CBP has E3/E4 ligase activity for p53.51),52) PCAF has E3 ligase activity toward Mdm253) and Gli1.54) The results of the present study demonstrated that the MYST domain, that is, the HAT domain, has an intrinsic ubiquitin E3 ligase activity. It is conceivable that other members of the MYST family HATs, such as MOF,55) MOZ/MORF,56) and yeast MYST proteins,57) have E3 ligase activity as well. Some functions of MYST family HATs might be attributable to E3 ligase activity, instead of HAT activity. For instance, Esa1 is essential for viability in budding yeast, but its HAT activity is dispensable for viability.58) The essential function of Esa1 may be its E3 ligase activity.
We thank Dr. Koji Okamoto for Mdm2 plasmid. M.I., M.A-T., T.S., and T.O. were supported by JSPS KAKENHI Grant Numbers JP26460400, JP15K08286, JP16K20163, and JP24591372, respectively.