2019 年 69 巻 2 号 p. 352-358
2019 年 69 巻 2 号 p. 352-358
IR64 is one of the world’s most popular rice cultivars. To collect genetic factors involved in controlling its heading date, we developed 70 reciprocal advanced-backcross populations with a total of 6284 individuals at the BC4F2 generation from crosses between Koshihikari and IR64. We detected 29 QTLs associated with heading date on chromosomes 3, 5–8, 10, and 12. Twenty QTLs were located in the same chromosome regions as previously isolated heading date genes (Hd1, Hd6, Hd16, Ghd7, DTH8, Hd17, and Hd18). The rest were located in other chromosome regions. We found more number of QTLs than previous studies using mapping populations of IR64. Fine mapping in additional advanced-backcross populations clearly revealed that QTLs on the long arm of chromosome 7 are overlapping and seem to be a novel genetic factor for heading date because of their different locations from OsPRR37. Our results suggest that the difference in heading date between IR64 and Koshihikari is genetically controlled by many factors, and that a non-functional allele of Hd1 contributes to early heading of IR64 in the genetic background of functional alleles of other heading date QTLs and genes such as Hd6 and Hd16.
Rice (Oryza sativa L.) is a staple food for more than half of the world’s population (Godfray et al. 2010). Heading date is one of the most important agronomic traits in rice, because it determines harvest seasons and planting regions. Modulation of heading date is effective for avoiding inappropriate environmental conditions for rice cultivation, such as times of drought, flood, and temperature extremes (Bentley et al. 2013). Changes in heading date can affect economic value through their effects on other agronomic traits such as yield and grain quality (Hori et al. 2012, Xue et al. 2008).
IR64 is an indica rice cultivar developed by the International Rice Research Institute in the Philippines in 1985 and is one of the world’s most popular cultivars (Mackill and Khush 2018), being widely grown in tropical and subtropical regions, including Southeast and South Asia, the Sahelian region of West Africa, and South America (Julia and Dingkuhn 2013, Khush and Virk 2005). It has high yield, good grain quality, and excellent palatability compared with other indica cultivars (Khush 1999, Khush and Virk 2005). By the end of the 20th century, it was grown on an estimated total of >10 million ha (Mackill and Khush 2018).
IR64 heads and matures early, with a total growth duration of about 117 days in the Philippines (Khush and Virk 2005). Genetic analyses of heading date in IR64 have reported up to 19 QTLs in doubled haploid lines derived from a cross between IR64 and a japonica rice cultivar, Azucena (Guo et al. 2013, Hittalmani et al. 2002, 2003, Li et al. 2003, Liu et al. 2007). Six QTLs were detected in backcrossed populations derived from crosses between IR64 and a wild relative, O. rufipogon IRGC 105491 (Septiningsih et al. 2003). Chromosomal locations of some of these QTLs coincided with genes that were isolated previously from cultivars with natural variation in heading date. Hd1 was the first heading date gene isolated from a QTL by map-based cloning (Yano et al. 2000). Sequencing analysis revealed that loss of a functional allele of Hd1 contributes to early heading date in IR64 (Itoh et al. 2018, Wei et al. 2016), which has functional alleles of most other heading date genes, such as Ghd7, DTH8, OsPRR37, Hd3a, RFT1, Hd6, and Hd16 (Itoh et al. 2018).
Advanced-backcross populations can reveal a large number of QTLs on various agronomic traits; both large- and small-effect QTLs were associated with the genetic architecture of agronomic traits in rice cultivars (Hori et al. 2015, Jing et al. 2010, Nagata et al. 2015, Xia et al. 2017). Here, to collect genetic factors involved in controlling heading date in IR64, we developed 6284 individual BC4F2 plants derived from reciprocal crosses between Koshihikari and IR64. QTL analysis revealed 20 QTLs that were located in regions of heading date genes isolated previously and 9 QTLs in other regions. The QTLs on the long arm of chromosome (Chr.) 7 were localized in a novel marker interval distinct from that of the heading date gene OsPRR37. These results will help our understanding of the genetic basis of the control of heading date in IR64.
We crossed Koshihikari and IR64 to obtain F1 individuals. We successively backcrossed progeny with either Koshihikari or IR64 to produce reciprocal BC4F1 plants (Supplemental Fig. 1). We selected 36 BC4F1 plants that had target heterozygous chromosome segments, minimizing the number of non-target chromosome segments from the donor (Supplemental Tables 1, 2). To detect heading date QTLs, we developed 31 BC4F2 populations in the Koshihikari background (K-01 to K-31) and 36 BC4F2 populations in the IR64 background (I-01 to I-36) by self-pollination of BC4F1 plants. We used three more BC4F2 populations (K-32, K-33, and I-37) and BC4F4 chromosome segment substitution lines (42 IRK-CSSLs and 40 KSI-CSSLs, Nagata et al. 2015) to confirm additive effects and chromosomal locations of the putative QTLs.
DNA extraction and genotyping by molecular markersTotal genomic DNA of individual plants was extracted from 1–3 cm of fresh leaves crushed in 250 μL extraction buffer containing 1 M KCl, 100 mM Tris·HCl (pH 8.0), and 10 mM EDTA (pH 8.0); precipitated with 100 μL 2-propanol; washed with 150 μL 70% ethanol; and dissolved in 30 μL buffer containing 1 mM Tris·HCl (pH 8.0) and 0.1 mM EDTA (pH 8.0) (Hori et al. 2015). We used simple sequence repeats (SSRs), insertion/deletion polymorphisms including gene-specific marker for linkage map construction and QTL detection. PCR amplification using 149 SSR markers and insertion/deletion markers for BX000503_3 and Hd1 and electrophoresis followed the method of Nagata et al. (2015).
Scoring of heading dateAll plants of the BC4F2 populations and the CSSLs were grown in an experimental field at the Institute of Crop Science, National Agriculture and Food Research Organization, Tsukuba, Japan (36.03°N, 140.11°E). Month-old seedlings were transplanted in mid May at one per hill in plots with a double row for each line at 18 cm between plants and 36 cm between rows. The mean day lengths were 13.1 h in April, 14.1 h in May, 14.6 h in June, 14.4 h in July, 13.5 h in August, and 12.4 h in September; floral transition occurs under natural long-day conditions around the summer solstice (Izawa 2007). Cultivation management followed the standard procedures used at the Institute. Days to heading of the individual plants were scored as the number of days from sowing to the appearance of the first panicle in 22 plants per population of K-01–31 and I-01–36, 186 plants per population of K-32, K-33, and I-37, and 5 plants per line of IRK-CSSLs and KSI-CSSLs.
Detection of heading date QTLsQTL analysis was performed by using composite interval mapping as implemented by the Zmapqtl program (model 6) provided by QTL Cartographer v. 2.5 software (Basten et al. 2005). Genome-wide threshold values (α = 0.05) were calculated from the results of 1000 permutations for QTL detection.
Average days to heading were 115.5 ± 2.6 in IR64 and 105.4 ± 1.5 in Koshihikari (Supplemental Tables 1, 2). To detect QTLs involved in heading date, we used 6284 plants at the BC4F2 generation derived from crosses between Koshihikari and IR64 (Supplemental Fig. 1). The BC4F2 populations showed a wide range of heading dates (Supplemental Tables 1, 2). Among 31 BC4F2 populations in the Koshihikari background, eight had segregating plants with an early heading date before Koshihikari of <102 days (K-10, K-14, K-15, K-16, K-21, K-24, K-25, and K-29), and one had segregating plants with a late heading date after IR64 of ≥124 days (K-09). Among 36 BC4F2 populations in the IR64 background, one had segregating plants with an early heading date of 105 days, the same as Koshihikari (I-06), and three had segregating plants with a late heading date after IR64 of >124 days (I-16, I-18, and I-19). The presence of several transgressive plants in BC4F2 populations with similar genetic backgrounds suggests that multiple heading date QTLs are involved in the difference between IR64 and Koshihikari.
QTL analyses in the BC4F2 populations detected 29 QTLs (Fig. 1, Table 1)—19 in the Koshihikari background and 10 in the IR64 background—of which 12 caused early heading with the IR64 allele and 17 with the Koshihikari allele. Three QTLs on the long arm of Chr. 6 showed large additive effects of >20 days in the IR64 background. One QTL on the long arm of Chr. 3 also showed a large additive effect of <–10 days in the Koshihikari background.
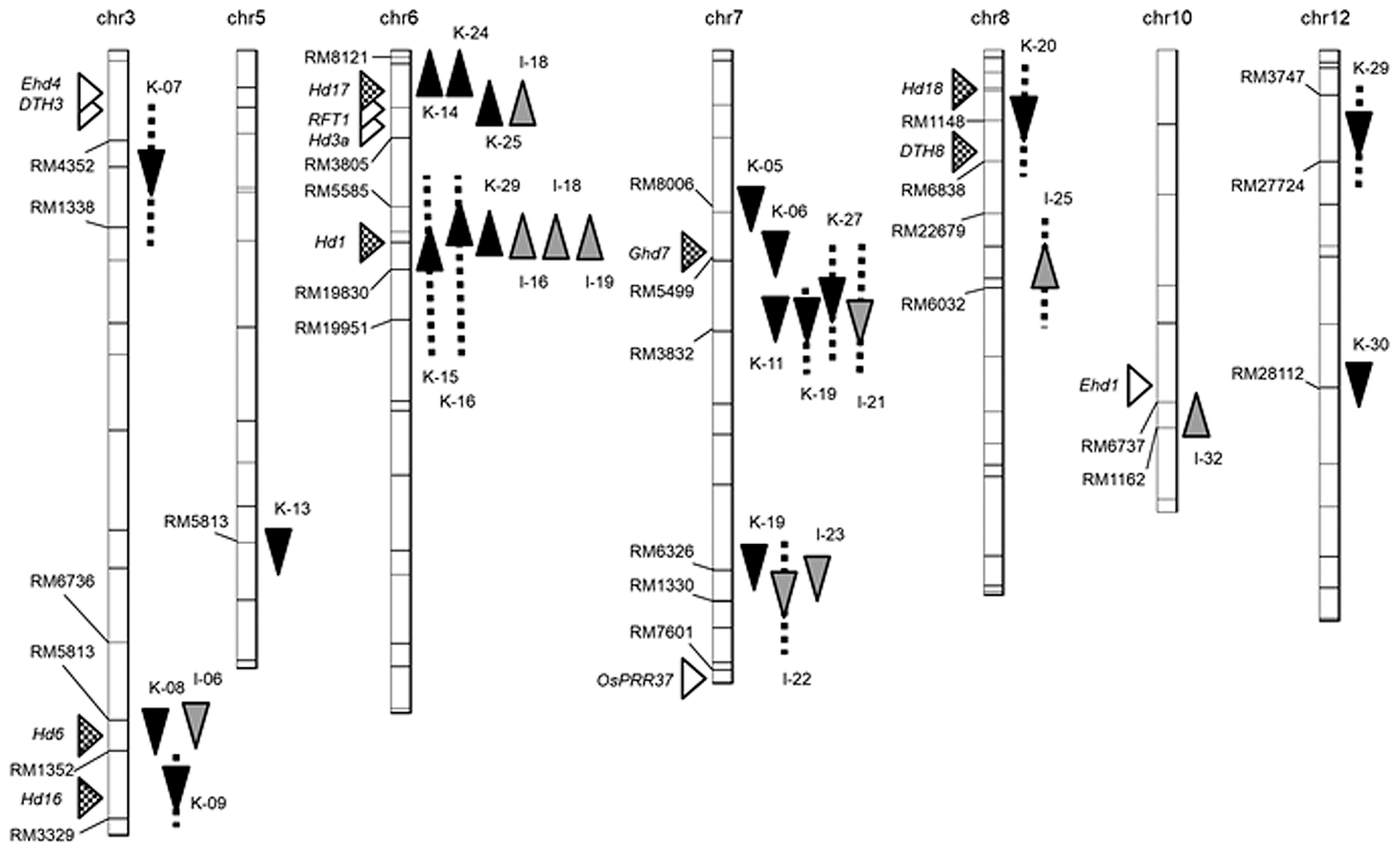
Chromosomal locations of QTLs for days to heading under natural field conditions. Arrowheads to the right of the linkage map indicate QTLs detected in BC4F2 populations with ▲▼ Koshihikari and

 IR64 genetic backgrounds: ▲
IR64 genetic backgrounds: ▲
 early and ▼
early and ▼
 late heading date by IR64 alleles. Triangles to the left indicate approximate locations of 13 heading date genes isolated previously.
late heading date by IR64 alleles. Triangles to the left indicate approximate locations of 13 heading date genes isolated previously.
 Heading date genes showing polymorphisms of the coding sequences between Koshihikari and IR64.
Heading date genes showing polymorphisms of the coding sequences between Koshihikari and IR64.
| A | ||||||||
|---|---|---|---|---|---|---|---|---|
| BC4F2 population | Chr. | Marker interval | Nearest marker | LODa | Additive effectb | Dominance effectc | PVE (%)d | Corresponding genee |
| K05 | 7 | RM7479–RM5499 | RM8006 | 5.4 | −4.4 | −1.2 | 64.0 | Ghd7 |
| K06 | 7 | RM8006–RM3832 | RM5499 | 5.7 | −5.0 | 0.6 | 67.0 | Ghd7 |
| K07 | 3 | RM4352–RM1338 | RM4352 | 5.6 | −4.1 | 1.2 | 76.0 | |
| K08 | 3 | RM6736–RM1352 | RM1352 | 7.1 | −4.1 | 0.3 | 56.0 | Hd6, Hd16 |
| K09 | 3 | RM1352–RM3329 | RM1352 | 16.3 | −12.6 | 5.1 | 92.0 | Hd6, Hd16 |
| K11 | 7 | RM5499–RM5481 | RM3832 | 4.1 | −3.2 | −0.7 | 55.0 | Ghd7 |
| K13 | 5 | RM4501–RM3809 | RM3476 | 3.7 | −2.3 | 0.3 | 52.0 | |
| K14 | 6 | RM8121–RM3805 | RM8121 | 2.8 | 3.1 | −0.8 | 44.0 | Hd17 |
| K15 | 6 | RM5585–RM19951 | RM5585 | 5.3 | 5.5 | −1.6 | 67.0 | Hd1 |
| K16 | 6 | RM5585–RM19951 | RM19951 | 8.4 | 6.1 | 0.6 | 85.0 | Hd1 |
| K19 | 7 | RM5499–RM3832 | RM5499 | 8.4 | −4.2 | 1.6 | 46.0 | Ghd7 |
| 7 | RM3404–RM1330 | RM6326 | 4.9 | −2.2 | −0.8 | 16.0 | ||
| K20 | 8 | RM1148–RM6838 | RM1148 | 8.0 | −4.6 | 1.0 | 83.0 | Hd18, DTH8 |
| K24 | 6 | RM8121–RM3805 | RM8121 | 5.9 | 2.8 | −0.3 | 68.0 | Hd17 |
| K25 | 6 | RM8121–RM4608 | RM3805 | 7.1 | 1.9 | 1.4 | 74.0 | Hd17 |
| K27 | 7 | RM8006–RM3832 | RM8006 | 8.1 | −5.0 | 1.1 | 67.0 | Ghd7 |
| K29 | 6 | Hd1ID006–RM19951 | RM19830 | 12.4 | 7.5 | 0.7 | 64.0 | Hd1 |
| 12 | RM3747–RM27724 | RM27724 | 3.9 | −2.8 | −0.6 | 6.0 | ||
| K30 | 12 | RM7102–RM28305 | RM28112 | 7.4 | −2.6 | −0.3 | 76.0 | |
| B | ||||||||
| BC4F2 population | Chr. | Marker interval | Nearest marker | LODa | Additive effectb | Dominance effectc | PVE (%)d | Corresponding genee |
| I06 | 3 | RM6736–RM1352 | RM5813 | 4.3 | −3.2 | 0.6 | 56.0 | Hd6, Hd16 |
| I16 | 6 | RM5585–RM19830 | Hd1ID006 | 23.1 | 37.7 | 21.8 | 93.7 | Hd1 |
| I18 | 6 | RM8121–RM4608 | RM8121 | 4.3 | 4.2 | 7.3 | 17.0 | Hd17 |
| 6 | RM5585–RM19830 | Hd1ID006 | 20.9 | 38.1 | 27.8 | 60.3 | Hd1 | |
| I19 | 6 | RM5585–RM19830 | Hd1ID006 | 29.7 | 38.4 | 21.6 | 95.0 | Hd1 |
| I21 | 7 | RM8006–RM3832 | RM8006 | 3.0 | −2.3 | 1.5 | 44.0 | Ghd7 |
| I22 | 7 | RM1330–RM7601 | RM1330 | 4.1 | −2.1 | −1.7 | 61.0 | |
| I23 | 7 | RM6326–RM1330 | RM6326 | 4.9 | −1.9 | −1.6 | 61.0 | |
| I25 | 8 | RM22679–RM6032 | RM6032 | 3.7 | 2.2 | −1.1 | 56.0 | |
| I32 | 10 | RM6737–RM1162 | RM1162 | 2.6 | 2.6 | 0.4 | 43.0 | |
The 29 QTLs were located on Chrs. 3, 5–8, 10, and 12. QTLs on the long arm of Chr. 6 would be the same as QTLs reported in previous studies using doubled haploid lines and backcrossed populations of IR64 (Guo et al. 2013, Hittalmani et al. 2002, 2003, Li et al. 2003, Liu et al. 2007, Septiningsih et al. 2003). QTLs on Chr. 3 were reported by Hittalmani et al. (2002 (2003) and Liu et al. (2007). QTLs on Chrs. 7, 10, and 12 were reported by Li et al. (2003), Liu et al. (2007), and Septiningsih et al. (2003). QTLs on Chr. 8 were reported by Hittalmani et al. (2002 2003), Li et al. (2003), Liu et al. (2007), and Septiningsih et al. (2003). However, our study using advanced-backcross populations found more QTLs than these previous studies using doubled haploid and backcrossed lines (2 to 19 QTLs). Advanced-backcross populations can detect small-additive-effect QTLs that are associated with the genetic architecture of various agronomic traits in rice cultivars, because the small-additive-effect QTLs were detected by separation from large-additive-effect QTLs (Hori et al. 2015, Jing et al. 2010, Nagata et al. 2015, Xia et al. 2017). Our study collected a lot of small-additive-effect QTLs involved in controlling heading date by using advanced-backcross populations of IR64 and Koshihikari.
We found 20 of the 29 QTLs near 7 heading date genes—Hd1, Hd6, Hd16, Ghd7, DTH8, Hd17, and Hd18—that show functional differences between IR64 and Koshihikari alleles. IR64 has a non-functional allele of Hd1 and functional alleles of other heading date genes (Ghd7, DTH8, OsPRR37, Hd3a, RFT1, Hd6, Hd16, Ehd1, Hd17, Ehd4, DTH3, and Hd18; Itoh et al. 2018, Shibaya et al. 2016, Wei et al. 2016). Koshihikari has non-functional alleles of Hd6 and Hd16 and functional alleles of the other genes. There are allelic differences in Ghd7, DTH8, Hd17, and Hd18 between IR64 and Koshihikari, although these genes are functional in both cultivars (Itoh et al. 2018, Matsubara et al. 2012, Shibaya et al. 2016).
Our results suggested that characteristics for heading date in IR64 consisted of a combination of large-additive-effect QTLs on the short arm of Chr. 6 (non-functional allele of Hd1) and the long arm of Chr. 3 (functional alleles of Hd6 and Hd16), and small-additive-effect QTLs in other regions. The genetic effect of Hd1 is larger in the IR64 background (with functional alleles of Hd6 and Hd16) than in the Koshihikari background (Supplemental Tables 1, 2; Nemoto et al. 2018, Ogiso et al. 2010). As indica rice cultivars, including IR64, have functional alleles of Hd6 and Hd16 (Hori et al. 2013, Yamane et al. 2009), the non-functional allele of Hd1 would be a major reason for the early heading and maturity of IR64 among indica rice cultivars. Thus, many genetic factors, including other small-additive-effect QTLs, contribute to the heading date of IR64.
Confirmation of a novel heading date QTL on the long arm of Chr. 7Among the 29 QTLs detected in the BC4F2 populations, we detected 3 QTLs on the long arm of Chr. 7 (Fig. 1, Table 1). These QTLs are overlapping but seem to lie at different marker intervals from OsPRR37, a key heading date gene localized at the distal end of the long arm of Chr. 7 (Koo et al. 2013). Both IR64 and Koshihikari have a functional allele of OsPRR37 (Itoh et al. 2018). Therefore, the QTLs detected in this region may be novel. To determine their precise chromosomal locations, we selected 368 BC4F2 plants in two populations in the Koshihikari background (K-32 and K-33) and 186 plants in one population in the IR64 background (I-37). K-32 had a heterozygous chromosome segment at RM1330 to the distal end of the long arm of Chr. 7, K-33 had a heterozygous segment at RM6326 to RM1330 (Supplemental Table 1), and I-37 had a heterozygous segment at RM6326 to the distal end of the long arm of Chr. 7 (Supplemental Table 2). Substitution mapping using the BC4F2 plants located these QTLs at the marker interval of RM6326 to RM7601, distinctly different from the genomic position of OsPRR37 (Fig. 2). We designated this novel QTL, at which the IR64 allele increases days to heading, as ‘qDTH7-3’, because dth7.2 has already been mapped on the long arm of Chr. 7 and qHD7.2 encodes OsPRR37 (Jing et al. 2018, Septiningsih et al. 2003).
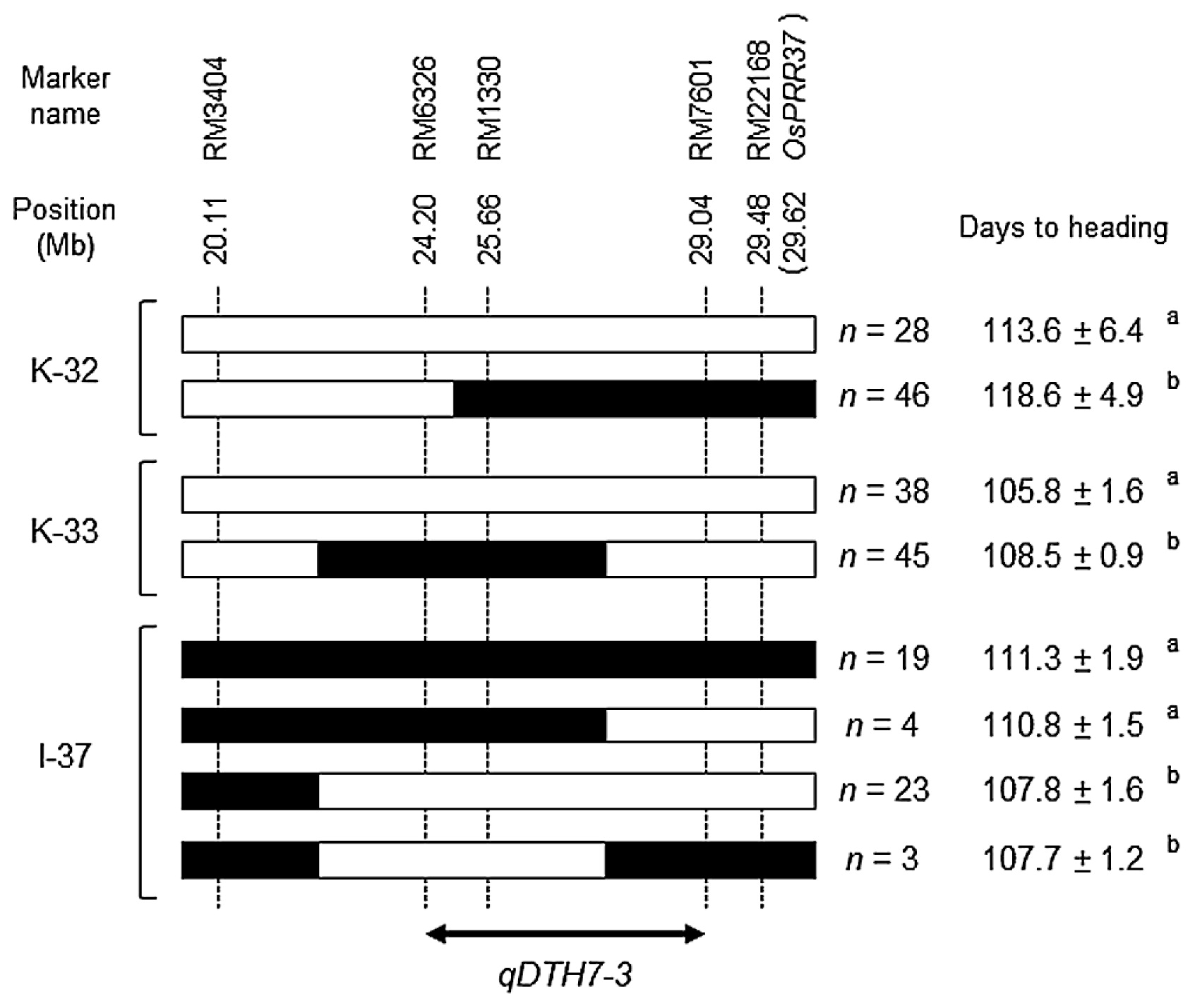
Graphical genotypes and positions of the QTLs for heading date that were detected on the long arm of Chr. 7 by the results of substitution mapping in three additional BC4F2 populations: 368 plants in the Koshihikari genetic background (K-32 and K-33) or 186 plants in the IR64 genetic background (I-37). ■ Homozygous for IR64, □ homozygous for Koshihikari. Significance levels calculated by the Tukey’s HSD test.
Nagata et al. (2015) developed a reciprocal set of CSSLs in the Koshihikari (42 IRK-CSSLs) and IR64 backgrounds (40 KSI-CSSLs) from the BC4F2 populations used here. Among the IRK-CSSLs, 14 lines had significant differences in heading date from Koshihikari (Supplemental Fig. 2). These lines have homozygous IR64 chromosome segments on the short and long arms of Chrs. 3, 6, 7, and 12 and on the short arm of Chr. 8. Among the KSI-CSSLs, 13 lines had significant differences in heading date from IR64. These lines have homozygous Koshihikari chromosome segments on the short and long arms of Chrs. 7 and 8, the long arms of Chrs. 3, 5 and 10, and the short arms of Chrs. 6 and 12. These chromosomal locations coincided well with those in the BC4F2 populations (Supplemental Fig. 2), supporting the existence of the heading date QTLs.
QTLs that determine the difference in heading date between IR64 and KoshihikariWe detected a number of QTLs involved in controlling heading date in IR64 and Koshihikari. At the genomic position of Hd1 on the long arm of Chr. 6, there were larger additive-effect QTLs in the BC4F2 populations and KSI-CSSLs of the IR64 background than in the BC4F2 populations and IRK-CSSLs of the Koshihikari background.
IR64 has a non-functional allele of Hd1 and functional alleles of other heading date genes. Koshihikari has non-functional alleles of Hd16 and Hd6 and functional alleles of Hd1 and other heading date genes. Thus, functional differences in Hd1, Hd16, and Hd6 would affect the phenotypic differences between the IR64 and Koshihikari backgrounds. IR64 is grown around the world in tropical and subtropical regions (Mackill and Khush 2018). The genetic effect of Hd1 is very strong in genetic backgrounds with functional alleles of Hd6 and Hd16 (Nemoto et al. 2018, Ogiso et al. 2010). Therefore, the non-functional allele of Hd1 is important to the early heading and maturity of indica rice cultivars in their cultivation areas. On the other hand, Koshihikari is grown throughout the temperate region in Japan (Hori et al. 2017, Kobayashi et al. 2018). The combination of the non-functional allele of Hd16 and the functional allele of Hd1 is a major reason for the early heading and maturity of Koshihikari in these region (Hori et al. 2013). However, Hd1 appears to be bifunctional in rice (Takahashi et al. 2009, Yano et al. 2000). Hd1 promotes flowering under short-day length conditions but represses flowering under long-day length conditions. And, there are complicated regulatory network of heading date genes in rice (Hori et al. 2016, Matsubara et al. 2014). In this study, we detected heading date QTLs by evaluating in the natural field (long-day length) conditions in the temperate region. Therefore, evaluation of the same advanced-backcross populations in different cultivation conditions (e.g. short-day length conditions in the tropical regions) would facilitate to reveal the overall genetic architecture involved in the control of heading date in IR64 and Koshihikari.
We also found six QTLs near Ghd7 on the long arm of Chr. 7 (Fig. 1, Table 1). Among them, QTLs detected in three populations in the Koshihikari background (K-11, K-19 and K-27) and one population in the IR64 background (I-21) might be novel, because these QTLs were localized on different marker intervals with the chromosomal position of Ghd7. And, one QTL was detected on the short arm of Chr. 8 including Hd18 and DTH8 in the Koshihikari background (K-20). Both IR64 and Koshihikari have functional alleles of Ghd7, Hd18 and DTH8, but there are non-synonymous substitutions in these heading date genes between the two rice cultivars (Itoh et al. 2018, Kumagai et al. 2013). Further analysis by using additional advanced-backcross populations is necessary to conclude whether these QTLs were novel or the same with one of either Ghd7, Hd18 or DHT8.
By using reciprocal advanced-backcross populations of BC4F2 plants, we revealed 29 QTLs, some near regions of previously isolated heading date genes and other in distinct regions. The QTLs on the long arm of Chr. 7 were localized in a novel marker interval different from OsPRR37. We also found difference of additive effects in the QTLs between the genetic backgrounds. However, to comprehensively elucidate genetic factors involved in controlling heading date in IR64, it is necessary to collect more QTLs by using other mapping populations derived from crosses between IR64 and many other cultivars. These results will help our understanding of the whole genetic structure involved in controlling heading date in IR64.
We are grateful to all staff of the Rice Applied Genomics Research Unit and the Field Management Division of NIAS/NARO for their technical assistance. This work was supported by the Ministry of Agriculture, Forestry and Fisheries of Japan (Integrated Research Project for Plant, Insect, and Animal using Genome Technology, QT1005; Genomics for Agricultural Innovation, NVR0001).