2019 Volume 42 Issue 3 Pages 432-441
2019 Volume 42 Issue 3 Pages 432-441
Moutan Cortex charcoal has been used to ameliorate blood heat symptoms and treat pathologic hemorrhage down the ages. Although well known as an agent with the effect of astringency and hemostasis, its active ingredients and action mechanism remain unclear. In the present study, molecular docking technology was employed to screen the potential hemostatic compounds in Moutan Cortex charcoal and their target proteins. Protein–protein-interaction (PPI) analysis was performed to explain the functions and enrichment pathways of the target proteins. The results showed that a total of 25 compounds were estimated as active constituents targeting multiple proteins related to hemostatic diseases, including 5 proteins (SERPINC1, FVIII, FX, FII and FXII) that were considered as the key targets. Then the drug-target (D-T) network was constructed to analyze the underlying hemostatic mechanism of Moutan Cortex charcoal, followed by a hierarchical cluster analysis (HCA) for compounds clustering, and a coagulation screening test for compound verification on their coagulation activities, with the results indicating that M15 (5-Tetradecenoic acid) and M31 (1-Monolinolein) might be the key compounds contributing to the hemostasis effect of Moutan Cortex charcoal by involving in the pathways related to complement, coagulation cascades and the platelet activation, particularly by activating FVIII, FX, FII and FXII and inhibiting SERPINC1. This study has demonstrated that Moutan Cortex charcoal may work as a hemostatic through the interaction between multiple-compounds and multiple-proteins, which provides the basis for further researches on the hemostasis mechanism of Moutan Cortex charcoal.
Medicinal carbon materials, as a popular hemostatic treatment, have been widely applied in clinical practice for over two thousand years. Moutan Cortex charcoal is the product of the root bark of Paeonia suffruticosa ANDREWS (Paeoniaceae), and it is produced after undergoing a processing step, in which the dried Moutan Cortex is stir-fried to be black-brown on the bark and burnt-brown inside.1) It has been used throughout the history of Traditional Chinese Medicine (TCM) from ancient China to modern society,2) with an early application dating back to the Yuan Dynasty in China as a component of the famous ancient prescription of “Shihui powder.”3) According to TCM theory, Moutan Cortex charcoal has the effect of blood cooling and hemostasis, and typically used for the treatment of uterine bleeding, gastrorrhagia, hematemesis, as well as other blood-heat and hemorrhage syndromes.1) Modern pharmacological investigations have also shown that Moutan Cortex charcoal is regarded as a satisfying hemostatic agent as demonstrated by significantly shortened bleeding time and clotting time in mice, as well as reduced rat plasma recalcification time (PRT), thrombin time (TT), prothrombin time (PT) and activated partial thromboplastin time (APTT). It can also increase the level of ADP and thromboxane B2 (TXB2) as well as collagen induced-platelet aggregation rate, in addition to a decrease in the content of 6-keto-prostaglandin F1α (PGF1α).4,5) To date, numerous researches have been conducted on Moutan Cortex charcoal, mainly focusing on the change of the chemical composition and its efficacy after processing.6–11) Moreover, the research strategy was basically routinized into chemical separation and analysis, components identification, and determination of the major active constituents, with the addition of some pharmacological experiments. However, the mechanism of its multi-compounds and multi-targets interaction remains to be revealed, let alone the low efficiency, big workload and cost of the separation and analysis work which is not uncommon to provide no benefit.
Network pharmacology has emerged as a new field of pharmacological study over the recent years.12) It aims to decipher the molecular mechanisms of the therapeutic effects of TCM and to determine their active ingredients or combinations.13–16) Estimating the potential key targets and pathways for the drugs against the diseases by computational methods is an efficient and time-saving approach for target discovery and experimental verification. Network pharmacology emphasizes “multi-components-multi-targets-multiple diseases” rather than “a drug-a gene-a disease” drug action pattern,17,18) which is becoming increasingly popular, particularly when deals with the complex systems.19–21) Researches based on network pharmacology for predicting the active ingredients and potential targets of traditional Chinese medicine have also been reported.22–28)
Therefore, we use network pharmacology approaches in this study, including molecular docking technology, protein–protein-interaction (PPI) analysis, drug-target (D-T) network, and hierarchical cluster analysis (HCA) in combination with a coagulation screening test to explore the potential pharmacodynamic material basis and the hemostasis mechanism of Moutan Cortex charcoal (procedure shown in Fig. 1 and the details are described in Table S1).

All the chemicals used in the experiments of this study were of reagent grade and obtained from commercial suppliers: Dade Actin Activated Cephaloplastin Reagent, Calcium Chloride Solution, Dade Fibrinogen Determination Reagents, Test Thrombin Reagent and Detergent for Fully Automated Blood Coagulation Analyzer (all from Siemens Healthcare Diagnostics Products GmbH, Germany); Disposable Vacuum Blood Vessels (SANLI, China); Chloral Hydrate (Damao Chemical Reagent Factory, Tianjin, China).
New Zealand white rabbits purchased from Laboratory Animal Center, Guangzhou University of Chinese Medicine (License No: SCXK (YUE) 2016-0041, Guangzhou, China); Compounds (M3, M9, M10, M11, M12, M15, M24, M31) were extracted, isolated and purified from Moutan Cortex Charcoal in our laboratory. The purity of the isolated components was shown to be higher than 96% by HPLC, and their structures were elucidated by comparing their spectroscopic data (electrospray ionization (ESI)-MS, 1H-NMR and 13C-NMR, heteronuclear multiple bond connectivity (HMBC), 1H-detected heteronuclear multiple quantum coherence (HMQC) and H–H correlation spectroscopy (COSY)) in literature.29,30) Structures of the eight compounds are shown in Fig. 2.

The compounds of Moutan Cortex charcoal were collected from the previous studies conducted by us and our peers.7,29–31) Advanced Chemistry Development (ACD/Labs) Software V11.0232) was used to predict compounds bioactivity, and compounds of Moutan Cortex charcoal were preliminarily screened according to “Lipinski’s rule of five” (Moriguchi octanol–water partition coeff (Log P ≤ 5), Molecular weight (MW ≤ 500), The number of acceptor atoms for H-bonds (HA ≤ 10), The number of donor atoms of H-bonds (HD ≤ 5), Freely rotatable bonds (FRB ≤ 10)).33) Molecules violating more than one of these rules may be inefficient in bioavailability and thus is considered unqualified for this study.
Target Protein ScreeningAs is known that blood coagulation mainly depends on the functioning of platelets34) and coagulation factors,35–37) the relevant proteins were collected from the protein structure database (PDB)38) to estimate the potential targets (Organism(s): Homo sapiens; Method: X-Ray diffraction; Resolution: > 1.5; Ligand: > 1) of Moutan Cortex charcoal for hemorrhagic diseases. Proteins were used to verify the reliability of the processing according to the following procedure (Fig. 3) by SYBYL-X 7.3 software (Tripos, U.S.A). First, the original ligand was pumped out, and then re-docked with the active pocket of the corresponding target, and total score (the total Surflex-Dock score expressed as −log(Kd)39)) was used as the threshold value. Root Mean Square Deviation (RMSD) between the conformation of re-docked ligand and the ligand in the original crystal structure was used to evaluate the reliability of the docking technology, and a RMSD value of less than 2 Å was deemed as an acceptable reliability for the docking method.40)

The other parameters are set to default, and are shown in Table S1.
The target proteins were imported to STRING to construct PPI network, with the species limited to “Homo sapiens” and a confidence score > 0.7.
Molecular DockingThe compounds were docked to target proteins using SYBYL-X 7.3 software. Based on the total score, potential active ingredients with an excellent binding to the target proteins were screened. The molecular docking process is shown in Fig. 3.
D-T Network ConstructionCytoscape is an open source software platform for biological pathways visualizing molecular interaction networks and integrating these networks with annotations, gene expression profiles, and other data.41–43) In order to provide a direct viewing on the binding between the active compounds and the target proteins, the results of molecular docking were visualized using Cytoscape software to construct the D-T network.
HCA of CompoundsBased on the data of “Lipinski’s rule of five,” compounds HCA was conducted using Ward Linkage method with SPSS 24.0.
Coagulation ScreeningTest Animal experiments of this Study were performed according to the Guide for the Care and Use of Laboratory Animals published by the National Institutes of Health (NIH Publications No. 85-23, revised 1996) and approved by the Ethics Committee of Guangdong Pharmaceutical University. New Zealand white rabbits (2.5 ± 0.2 kg) were housed in accordance with guidelines on Laboratory Animal Requirements of Environment as well as the requirements from the Committee on Ethics of Animal Experiments. The rabbits, after being fasted for 8 h, were anesthetized with 10% chloral hydrate. The blood was drawn from the heart and collected in a plastic centrifuge tube containing 3.8% sodium citrate (volume ratio 9 : 1) as the anticoagulant, followed by a 10-min 3000 r/min centrifugation to separate the plasma for further analysis.44) Activated partial thromboplastin time (APTT), thrombin time (TT) and fibrinogen content (FIB) were used as the coagulation indexes. The monomer compounds were respectively dissolved in methanol to prepare solutions of 0.03 g/mL, with methanol solvent used as blank control. Nine hundred microliters of plasma was transferred into centrifuge tube, and then 10 µL monomer solution was added for determination by coagulation method according to the instructions of the kits. The results of coagulation screening test were expressed as means ± standard deviation (S.D.). The significance of inter-group differences was analyzed by one-way ANOVA using SPSS 24.0 statistical software. p-value of less than 0.05 or 0.01 was considered statistically significant.
A total of 40 compounds, including 5 compounds identified by the previous research and another 35 that were separated and authenticated by our lab, were collected for this study (Compounds information was reported in another paper). The bioactivity parameters of these compounds under “Lipinski’s rule of five” were obtained with Chemistry Development (ACD/Labs) Software V11.02 (Table 1), and each of the 40 compounds followed more than three of the five rules.
| Name | Compound name | CAS | Log P | MW | HA | HD | FRB |
|---|---|---|---|---|---|---|---|
| M1 | Paeonol | 552-41-0 | 2.309 | 166.17 | 3 | 1 | 3 |
| M2 | 3-(2-Furanyl)-1-(2-hydroxy-4-methoxyphenyl)-2-propen-1-one | 52542-11-7 | 3.369 | 244.24 | 4 | 1 | 5 |
| M3 | Dibutyl phthalate | 84-74-2 | 4.752 | 278.34 | 4 | 0 | 10 |
| M4 | 3,6-Dimethyl-5-benzofuranol | 59211-24-4 | 3.224 | 162.19 | 2 | 1 | 1 |
| M5 | Diethyl phthalate | 84-66-2 | 2.714 | 222.24 | 4 | 0 | 6 |
| M6 | α1,α2-Dethyl-1,2-benzenedimethanol | 196871-46-2 | 1.616 | 194.27 | 2 | 2 | 6 |
| M7 | Betaprost | 83-46-5 | 10.482 | 414.72 | 1 | 1 | 7 |
| M8 | Benzoic acid | 65-85-0 | 1.559 | 122.12 | 2 | 1 | 1 |
| M9 | 9-Octadecenoic acid-2-hydroxyethyl ester | 25905-73-1 | 7.219 | 326.51 | 3 | 1 | 19 |
| M10 | Methyl tridecanoate | 1731-88-0 | 5.88 | 228.37 | 2 | 0 | 12 |
| M11 | 1-Monomyristin | 589-68-4 | 5.048 | 302.45 | 4 | 2 | 18 |
| M12 | 3-(Decyloxy)-1-propanol | 60851-88-9 | 4.302 | 216.37 | 2 | 1 | 13 |
| M13 | 1,4-Diethyl-cyclohexane | 10355-58-5 | 6.272 | 168.32 | 0 | 0 | 4 |
| M14 | 1-(3,5-Dihydroxy-4-methylphenyl)-ethanone | 855925-40-5 | 1.101 | 166.17 | 3 | 2 | 3 |
| M15 | Methyl physeterate | 103385-67-7 | 5.965 | 240.38 | 2 | 0 | 12 |
| M16 | Betulinic acid | 472-15-1 | 7.653 | 456.7 | 3 | 2 | 3 |
| M17 | Oleanolic acid | 508-02-1 | 8.576 | 456.7 | 3 | 2 | 2 |
| M18 | 1-(2,4-Dihydroxyphenyl)-ethanone | 74291-78-4 | 1.48 | 152.15 | 3 | 2 | 3 |
| M19 | Anisic acid | 100-09-4 | 1.776 | 152.15 | 3 | 1 | 2 |
| M20 | 1-(2,5-Dihydroxy-4-methoxyphenyl)-1-propanone | 3839-58-5 | 2.059 | 196.2 | 4 | 2 | 5 |
| M21 | 1-(2,5-Dihydroxy-4-methylphenyl)-ethanone | 54698-17-8 | 1.438 | 166.17 | 3 | 2 | 3 |
| M22 | Oxyphenic acid | 120-80-9 | 0.844 | 110.11 | 2 | 2 | 2 |
| M23 | 1-(2,5-Dihydroxy-4-methoxyphenyl)-ethanone | 22089-12-9 | 1.549 | 182.17 | 4 | 2 | 4 |
| M24 | 9,12-Octadecadienoic acid methyl ester | 2462-85-3 | 7.615 | 294.47 | 2 | 0 | 15 |
| M25 | Paridol | 99-76-3 | 1.882 | 152.15 | 3 | 1 | 3 |
| M26 | 4-Acetylresorcinol | 89-84-9 | 1.48 | 152.15 | 3 | 2 | 3 |
| M27 | 2,3-Dihydroxy-4-methoxyacetophenone | 708-53-2 | 1.773 | 182.17 | 4 | 2 | 4 |
| M28 | Larixic acid | 118-71-8 | 0.07 | 126.11 | 3 | 1 | 1 |
| M29 | Alginetin | 6005-10-3 | 1.32 | 192.17 | 4 | 2 | 2 |
| M30 | β-Amyrenol | 559-70-6 | 10.481 | 426.72 | 1 | 1 | 1 |
| M31 | 1-Monolinolein | 2277-28-3 | 6.273 | 354.52 | 4 | 2 | 20 |
| M32 | 2,5-Dihydroxy-α,4-dimethyl-benzeneacetic acid ethyl ester | 2114236-13-2 | 1.641 | 224.25 | 4 | 2 | 6 |
| M33 | 5-Acetyl-2-methoxyphenol | 6100-74-9 | 1.373 | 166.17 | 3 | 1 | 3 |
| M34 | Betulinic acid methyl ester | 2259-06-5 | 9.193 | 470.73 | 3 | 1 | 4 |
| M35 | Paeoniflorin | 23180-57-6 | 0.245 | 480.46 | 11 | 5 | 12 |
| M36 | Quercetin | 117-39-5 | 1.989 | 302.24 | 7 | 5 | 6 |
| M37 | Kaempferid | 491-54-3 | 3.499 | 300.26 | 6 | 3 | 5 |
| M38 | Gallic acid | 149-91-7 | 0.531 | 170.12 | 5 | 4 | 4 |
| M39 | Isorhamnetol | 480-19-3 | 2.787 | 316.26 | 7 | 4 | 6 |
| M40 | 5-Hydroxymethylfurfural | 67-47-0 | −0.778 | 126.11 | 3 | 1 | 3 |
Following the procedure for coagulation protein screening as shown in Fig. 3, 15 proteins were eventually estimated as the target proteins according to the criteria of RMSD < 2 Å. The total score and RMSD values of these target proteins are shown in Table 2.
| Target | Abbreviation | PDB-ID | Total score | RMSD(Å) |
|---|---|---|---|---|
| Urokinase-type plasminogen activator | PLAU | 1EJN | 8.92 | 1.51 |
| Ras-related protein Rap-1A | RaP1A | 4KVG | 8.68 | 1.70 |
| Activated protein C | PROC | 1AUT | 7.34 | 1.79 |
| Coagulation factor XI | FXI | 4D76 | 7.23 | 1.69 |
| Coagulation factor IX | FIX | 5JBC | 6.37 | 1.51 |
| Plasma kallikrein | KLKB1 | 2ANY | 6.17 | 0.14 |
| Platelet glycoprotein Ib alpha chain | GP1BA | 1QYY | 6.14 | 0.78 |
| Coagulation factor VII | FVII | 4ZXY | 5.77 | 1.54 |
| Coagulation factor XII | FXII | 4XE4 | 5.40 | 1.03 |
| Guanine nucleotide-binding protein G(s) subunit alpha | GNAS | 5G53 | 5.32 | 1.84 |
| Coagulation factor VIII | FVIII | 3J2Q | 5.28 | 1.05 |
| Antithrombin-III | SERPINC1/AT-III | 2ZNH | 5.28 | 0.67 |
| Fibrin beta chain | FGB | 1N86 | 5.24 | 0.70 |
| Coagulation factor X | FX | 1G2M | 5.11 | 0.82 |
| Prothrombin | FII | 3TU7 | 5.11 | 1.45 |
The PPI network of the target proteins was constructed, as illustrated in Fig. 4. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis reveals that the 15 proteins are mainly enriched in pathways related to complement, coagulation cascades and platelet activation, and more than 10 out of the 15 proteins involve in the biological process of blood coagulation and fibrin clot formation according to Gene Ontology (GO) analysis (Table 3).

(Color figure can be accessed in the online version.)
| Term | ID | Description | Count | FRD |
|---|---|---|---|---|
| Biological process (GO) | GO:0072378 | Blood coagulation, fibrin clot formation | 10 | 3.82E-23 |
| GO:0007596 | Blood coagulation | 15 | 2.44E-21 | |
| Molecular function (GO) | GO:0004252 | Serine-type endopeptidase activity | 9 | 1.29E-13 |
| GO:0016787 | Hydrolase activity | 10 | 1.57E-04 | |
| Cellular component (GO) | GO:0005615 | Extracellular space | 11 | 4.60E-08 |
| GO:0005796 | Golgi lumen | 5 | 1.75E-06 | |
| KEGG pathways | 04610 | Complement and coagulation cascades | 12 | 8.44E-26 |
| 04611 | Platelet activation | 4 | 2.55E-04 |
A total of 40 compounds were selected for docking to the 15 target proteins under the procedure shown in Fig. 3, and the molecular docking scores of the compounds are summarized in Table 4. The total score of the known small molecular ligand docking on the target protein complex was employed as the threshold, so the compounds docking on this protein with a total score greater than this threshold value is considered as the main compounds acting on the target.45) The results revealed that 25 out of the 40 compounds interacted with the 15 hemostasis target proteins (Table 4).
| Name | PLAU | RaP1A | PROC | FXI | FIX | KLKB1 | GP1BA | FVII | FXII | GNAS | FVIII | AT-III | FGB | FX | FII |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| M1 | — | — | — | — | — | — | — | — | — | — | — | — | — | 5.67 | — |
| M2 | — | — | — | — | — | — | 6.64 | — | — | — | — | 5.64 | — | — | — |
| M3 | — | — | — | — | — | 6.32 | — | — | 5.54 | 6.14 | 7.41 | 5.56 | 5.65 | 5.27 | 6.71 |
| M4 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | — |
| M5 | — | — | — | — | — | — | — | — | 5.41 | — | — | — | — | — | — |
| M6 | — | — | — | — | — | — | — | — | — | — | 5.37 | — | — | — | — |
| M7 | — | — | — | — | 6.50 | — | — | — | — | 5.77 | 7.72 | 6.81 | — | 5.85 | 5.48 |
| M8 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | — |
| M9 | — | — | — | — | 7.63 | 6.17 | — | 7.42 | 9.88 | 8.65 | 9.75 | 8.13 | 9.60 | 7.31 | 7.29 |
| M10 | — | — | — | — | — | — | 5.76 | — | 7.01 | 6.00 | 5.49 | 6.83 | 6.31 | 5.97 | 7.23 |
| M11 | — | — | — | 7.37 | — | 5.40 | 7.80 | 7.52 | 7.76 | 7.45 | 7.67 | 8.39 | 8.15 | 6.47 | 6.87 |
| M12 | — | — | — | — | — | 6.41 | — | 6.72 | 7.18 | 6.43 | 5.98 | 7.03 | 5.38 | 5.32 | 6.96 |
| M13 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | — |
| M14 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | — |
| M15 | — | — | — | — | — | — | 6.27 | 5.85 | 7.02 | 6.34 | 7.57 | 6.45 | 5.83 | 6.26 | 7.21 |
| M16 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | — |
| M17 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | — |
| M18 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | — |
| M19 | — | — | — | — | — | 6.95 | — | — | 5.44 | — | 6.35 | 5.44 | — | — | 5.49 |
| M20 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | — |
| M21 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | — |
| M22 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | — |
| M23 | — | — | — | — | — | — | — | — | — | — | — | 6.01 | — | — | — |
| M24 | — | — | — | 7.71 | 6.55 | — | 7.50 | 5.83 | 8.49 | 6.33 | 8.68 | 8.93 | 7.10 | 5.49 | 8.14 |
| M25 | — | — | — | — | — | — | — | — | 9.43 | 7.19 | — | — | — | — | — |
| M26 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | — |
| M27 | — | — | — | — | — | — | — | — | — | — | — | — | 5.24 | 5.54 | — |
| M28 | — | — | — | — | — | — | 6.90 | — | — | — | — | — | — | — | — |
| M29 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | — |
| M30 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | — |
| M31 | — | — | — | — | — | 7.17 | 8.49 | 9.74 | 8.36 | 8.10 | 8.25 | 8.93 | 8.80 | 8.07 | 6.66 |
| M32 | — | — | — | — | — | — | — | 6.87 | 7.83 | — | — | 7.48 | — | — | 5.24 |
| M33 | — | — | — | — | — | — | — | — | — | — | — | — | — | 5.14 | — |
| M34 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | — |
| M35 | — | — | — | — | — | 6.18 | 6.50 | 6.01 | — | — | 5.69 | 7.25 | — | — | — |
| M36 | — | — | — | — | — | — | — | — | 6.10 | — | 5.99 | — | — | — | — |
| M37 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | 5.47 |
| M38 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | 5.13 |
| M39 | — | — | — | — | — | — | — | 5.79 | — | 7.11 | 8.49 | — | 5.38 | 6.32 | — |
| M40 | — | — | — | — | — | — | — | — | — | — | — | — | — | — | — |
D-T network was constructed, as shown in Fig. 5. Betweenness, degree and closeness were employed as the parameters to assess the binding activity of the compounds as well as the influence of the target proteins on the whole network (Tables 5, 6). Protein with a higher score reflects a greater influence, and can be regarded as a key protein in the network. Analysis of the D-T network revealed that 8 compounds (M3, M9, M10, M11, M12, M15, M24 and M31) demonstrated a good binding activity with 8 or more target proteins (Betweenness > 0.032, Closeness > 0.51, degree ≥ 8, Fig. 5, Table 5), and five proteins (SERPINC1, FVIII, FX, FII and FXII) qualified as the key ones related to the hemostatic effect of Moutan Cortex charcoal (Betweenness > 0.129, Closeness > 0.50, degree ≥ 13, Fig. 5, Table 6).

The edges represent interactions between the compounds and the targets. The network was constructed and visualized with Cytoscape. (Color figure can be accessed in the online version.)
| Name | Betweenness | Closeness | Degree | Name | Betweenness | Closeness | Degree |
|---|---|---|---|---|---|---|---|
| M24 | 0.0967 | 0.5806 | 11 | M2 | 0.0015 | 0.3600 | 2 |
| M11 | 0.0852 | 0.5806 | 11 | M25 | 0.0007 | 0.3600 | 2 |
| M31 | 0.0579 | 0.5625 | 10 | M27 | 0.0005 | 0.3462 | 2 |
| M9 | 0.0585 | 0.5455 | 10 | M36 | 0.0011 | 0.3750 | 2 |
| M15 | 0.0487 | 0.5455 | 9 | M1 | 0.0000 | 0.3396 | 1 |
| M12 | 0.0392 | 0.5294 | 9 | M23 | 0.0000 | 0.3462 | 1 |
| M10 | 0.0417 | 0.5294 | 8 | M28 | 0.0000 | 0.3103 | 1 |
| M3 | 0.0328 | 0.5143 | 8 | M33 | 0.0000 | 0.3396 | 1 |
| M7 | 0.0243 | 0.4737 | 6 | M37 | 0.0000 | 0.3396 | 1 |
| M19 | 0.0118 | 0.4390 | 5 | M38 | 0.0000 | 0.3396 | 1 |
| M35 | 0.0113 | 0.4091 | 5 | M5 | 0.0000 | 0.3396 | 1 |
| M39 | 0.0081 | 0.4186 | 5 | M6 | 0.0000 | 0.3462 | 1 |
| M32 | 0.0070 | 0.4186 | 4 |
| Name | Betweenness | Closeness | Degree | Name | Betweenness | Closeness | Degree |
|---|---|---|---|---|---|---|---|
| SERPINC1 | 0.1351 | 0.5217 | 14 | FGB | 0.0398 | 0.4675 | 10 |
| FVIII | 0.1264 | 0.5217 | 14 | FVII | 0.0270 | 0.4557 | 9 |
| FX | 0.1582 | 0.5070 | 13 | GP1BA | 0.0795 | 0.4444 | 8 |
| FII | 0.1414 | 0.5070 | 13 | KLKB1 | 0.0126 | 0.4337 | 7 |
| FXII | 0.1291 | 0.5070 | 13 | FIX | 0.0015 | 0.3956 | 3 |
| GNAS | 0.0475 | 0.4800 | 11 | FXI | 0.0003 | 0.3871 | 2 |
Forty compounds were classified into four groups, as demonstrated in Fig. 6, and the results of HCA Analysis is shown in Table 7. Significant inter-group difference was observed in MW value among the four groups, as contrasted to HA and HD values in which the paired inter-group comparisons between any two of the four groups showed no remarkable difference. As for Log P and FRB, statistical difference was found in most of the paired-comparisons, except for the second group and the third group, which showed no difference in either of the two parameters. This may explain why the second group and the third group are closer to each other in the HCA chart, while the first group and the fourth group are farther apart. Moreover, 8 compounds (M3, M9, M10, M11, M12, M15, M24 and M31) with strong binding activity all fell in the second and the third groups, and those (M4, M8, M13, M14, M16, M17, M18, M20, M21, M22, M26, M29, M30, M34, M40) with no binding activity all clustered in the first and the fourth groups. Combining with the molecular docking results, we may conclude that the first and the fourth groups have the compounds with low or no binding activity, while those in the second and the third groups are likely to have a stronger binding activity with the target proteins.
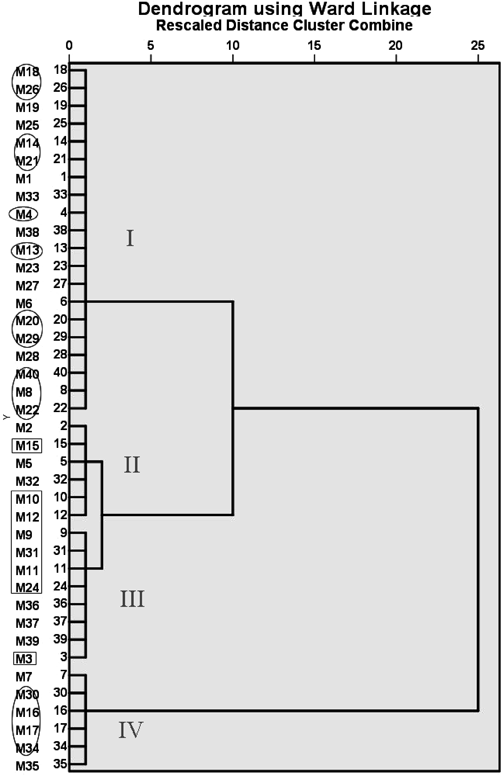
Ellipses represent inactive compounds estimated through virtual screening on the results of molecular docking. Diamonds represent the 8 compounds with the highest scores in Cytoscape analysis.
| Average | Log P | MW | HA | HD | FRB |
|---|---|---|---|---|---|
| First group | 1.64*☆☆△△ | 160.27**☆☆△△ | 3☆ | 2 | 3**☆☆ |
| Second group | 3.98▲ | 229.31○○▲▲ | 3 | 1○ | 9 |
| Third group | 4.90● | 309.38●● | 5 | 2 | 12●● |
| Fourth group | 7.77 | 451.01 | 4 | 2 | 5 |
** p < 0.01, * p < 0.05 (First group vs. Second group); ☆☆ p < 0.01, ☆ p < 0.05 (First group vs. Third group); △△ p < 0.01, △ p < 0.05 (First group vs. Fourth group); ○○ p < 0.01, ○ p < 0.05 (Second group vs. Third group); ▲▲ p < 0.01, ▲ p < 0.05 (Second group vs. Fourth group); ●● p < 0.01, ● p < 0.05 (Third group vs. Fourth group).
As eight compounds (M3, M9, M10, M11, M12, M15, M24 and M31) were preliminarily predicated by D-T network having a strong binding activity with the target proteins, coagulation screening test for them were performed in vitro, using APTT, FIB and TT as the indexes for analysis (Table 8). The APTT mainly reflects the activity and content of endogenous coagulation factors, and TT is a parameter indicating the coagulation, anticoagulation and fibrinolysis system, while FIB words as the raw material of synthetic fibrin, and plays a significant role in coagulation process.46) As compared with the normal group, APTT was significantly increased by compound M9, M10 and M12, and the level of FIB was dramatically decreased by compound M12 but showed a remarkable increase in compound M15 and M31 groups (p < 0.01). The results suggest that compound M15 and M31 are likely to have a coagulation effect, while compound M9, M10 and M12 appear to demonstrate an anticoagulation effect. These inferences on their biological activity need further verification in more in vivo studies. In this study, we found that the processing of Moutan Cortex led to a decrease of M3, M11 and M24 as well as an increase of M9, M10 and M12 in Moutan Cortex charcoal. Also, other researches reported an elevated level of hemostatic components (gallic acid and 5-hydroxymethylfurfural (5-HMF)) in contrast to a significant decline in blood activating components (quercetin, kaempferol and isorhamnetin) after the stir-fried processing of Moutan Cortex.6–9,47–50) According to the theory in traditional Chinese Medicine (TCM), Moutan Cortex has a function of promoting blood circulation, and the carbonizing process can weaken its blood activating while at the same time enhance its hemostasis effect. Moutan Cortex charcoal is therefore regarded as a hemostatic without the consequence of blood stasis. Based on these findings and the theory, we conclude that among the five active constituents (M15, M31, M9, M10 and M12, Fig. 2), compounds M15 and M31 are most likely to be the key ones responsible for the hemostasis effect of Moutan Cortex charcoal.
| No. | C (g/mL) | APTT (s) | FIB (g/L) | TT (s) | No. | C (g/mL) | APTT (s) | FIB (g/L) | TT (s) |
|---|---|---|---|---|---|---|---|---|---|
| Normal | — | 21.8 | 1.77 | 19.7 | Normal | — | 21.8 | 1.77 | 19.7 |
| M3 | 0.003 | 27.5** | 1.90* | 19.3 | M12 | 0.003 | 29.6** | 1.62* | 18.9 |
| M9 | 0.003 | 29.1** | 1.79 | 19.6 | M15 | 0.003 | 24.9 | 1.93** | 19.9 |
| M10 | 0.003 | 30.5** | 1.75 | 21.1** | M24 | 0.003 | 29.5** | 2.00** | 19.3 |
| M11 | 0.003 | 24.1 | 1.86 | 19.3 | M31 | 0.003 | 19.4 | 2.34** | 19.3 |
* p < 0.05, ** p < 0.01 vs. normal group.
The results delineated as above can lead that compounds M15 and M31 are the hemostasis active components, and close binding to 5 target proteins (SERPINC1, FVIII, FX, FII and FXII). Factor VIII is important for the generation of amplified factor Xa and sustained coagulation.51) FX can be converted to activated FX (FXa) via either the intrinsic or extrinsic pathway, which, by interacting with activated factor V, can activate prothrombin (FII), eventually triggering the coagulation cascade.52) AT-III (SERPINC1), as the most important blocking factor of active blood coagulation factors, can regulate blood coagulation and fibrinolysis.53) And Factor XII (FXII) is one of the elements involving in the contact activation coagulation system by working together with prekallikrein and the cofactor of high molecular weight kininogen.54) So presumably compound M15 and M31 collected from Moutan Cortex charcoal might play a role in coagulation by activating FVIII, FX, FII and FXII, and in the meantime, inhibiting the activity of SERPINC1.
In this study, 40 active compounds of Moutan Cortex charcoal were obtained by rapid screening and then semiflexibly docked to 15 proteins involved in hemostasis. The results reveal that Moutan Cortex charcoal may act as a blood coagulation promoter mainly through the interaction between two compounds (M15 and M31) and five target proteins (SERPINC1, FVIII, FX, FII and FXII) which are involved in the pathways associated with complement and coagulation cascades as well as platelet activation. Moreover, M15 and M31 possibly play their parts by a positive effect on FVIII, FX, FII and FXII as well as an inhibitory action on SERPINC1. This study provides a basis for further exploration on the hemostasis mechanism of Moutan Cortex charcoal using a multi-compounds and multi-targets approach. Plus, it also provides a reference for the studies on the hemostasis mechanism of Chinese charcoal medicines.
The authors sincerely thank all the volunteers for their participation in this study. This work was supported by the National Natural Sciences Foundation of China (No. 81473352); the Guangdong Province innovation training project (No. 201610573040); the Guangzhou science and technology planning project (No. 201707010170).
The authors declare no conflict of interest.
The online version of this article contains supplementary materials.