2019 Volume 44 Issue 6 Pages 425-433
2019 Volume 44 Issue 6 Pages 425-433
Cardiac fibroblasts (CFs) could be activated after myocardial infarction (MI). Thus, it is necessary to explore effective drugs to suppress the activation of CFs following MI. This study was designed to investigate the impacts of ellagic acid on CFs and the underlying mechanisms. The expression of histone deacetylases (HDACs) and fibrosis-related genes was detected by qRT-PCR and western blot. The Masson’s Trichrome Staining assay was used to evaluate the area of cardiac fibrosis. The proliferation and migration of CFs were measured by CCK8 Kit and Transwell assay, respectively. Our results showed that ellagic acid significantly reduced protein expression of HDAC1, mRNA expression of collagen I, collagen III, MMP-2 and MMP-9 and the area of cardiac fibrosis in MI rats. In Ang II-stimulated CFs, ellagic acid (60 μmol/L) decreased the protein expression of HDAC1, collagen I, collagen III, MMP-2 and MMP-9, and inhibited cell proliferation and migration. Further, HDAC1 over-expression reversed the inhibitor effects of ellagic acid on proteins expression (collagen I, collagen III, MMP-2 and MMP-9) and proliferation and migration of CFs. The present results suggested that ellagic acid suppressed proliferation and migration of CFs by down-regulating expression of HDAC1.
Myocardial infarction (MI) is the primary reason of heart failure, which leads to high morbidity and mortality (Tamargo and López-Sendón, 2011; Redfield, 2002). Cardiac fibroblasts (CFs) are activated during the development of heart failure caused by MI. Then, the CFs proliferate and secrete lots of proteins including collagens. Collagens compose the extracellular matrix (ECM), and replace the necrotic myocardial cells to maintain cardiac structure (Sui et al., 2015). Nevertheless, over-activity of CFs results in excessive accumulation of ECM, induces cardiac fibrosis (Fan et al., 2012), and finally causes myocardial elasticity impairment and heart failure (Zamilpa and Lindsey, 2010). Therefore, it is necessary to explore the effective medicine to suppress the activation of CFs caused by MI.
Ellagic acid, a dimer derivative of gallic acid with trans-gallic acid tannin structure, is one of the major phenolic substances in pomegranate (Landete, 2011). Recent studies observed the vital function of ellagic acid in protecting cardiovascular system. For example, ellagic acid could reduce the area of MI and suppress cardiac fibrosis by regulating the expression of anti-apoptosis genes and enhancing activities of mitochondrial respiratory enzymes (Mari Kannan and Darlin Quine, 2012; Wei et al., 2017). Furthermore, the occurrence of ventricular hypertrophy could be decreased by ellagic acid via the suppression of lipid peroxidation (Kannan and Quine, 2013). However, there were scarce studies in regards to the effects of ellagic acid on CFs and the underlying mechanism.
As important epigenetic regulatory enzymes, histone deacetylases (HDACs) can stimulate cardiac hypertrophy and cardiac fibrosis. Previous studies revealed that inhibitors of HDACs relieved cardiac hypertrophy pathologically (Iyer et al., 2010; Cardinale et al., 2010; Kee et al., 2006; Kook et al., 2003), and attenuate the myocardial remodeling following MI (Lee et al., 2007). More importantly, a study showed that the proliferation and migration of CFs have been suppressed by an inhibitor of HDACs (Kao et al., 2013). Another study also revealed that inhibition of HDACs could repress cardiac fibrosis mediated by Ang II via CFs and fibroblasts derived from bone marrow (Williams et al., 2014). In our preliminary experiments, we found that ellagic acid could affect the expression of HDAC1, while the expression of HDAC2 and HDAC6 had no alterations.
According to the functions of HDACs in cardiac fibrosis and the effect of ellagic acid on the expression of HDAC1, we hypothesized that ellagic acid might inhibit proliferation and migration of CFs through down-regulation expression of HDAC1. In the present study, we aim to verify our hypothesis and elucidate the mechanism underlying the effects of ellagic acid on CFs. In vivo, we detected expressions of HDACs and fibrosis-related factors, and observed the alterations of fibrosis area in rat model of MI. In vitro, the expressions of HDAC1 and fibrosis-related factors were measured, the proliferation and migration of CFs were evaluated, and the fibrosis area was observed.
Male SD rats (8 weeks old, 200-230 g) were obtained from the Animal Experimental Center, Guangdong Academy of Medical Science (Guangdong, China). Animals were raised in standard pathogen-free conditions. The illumination cycle was 12 hr light/dark, the temperature was 20-25°C and the humidity in atmosphere was 55%-60%. All of them were kept with free access to food and water and allowed to acclimatize for 1 week before starting the formal experiment. All experimental procedures performed were approved by the Committee on Animal Care of The Second Affiliated Hospital and Yuying Children’s Hospital of Wenzhou Medical University.
Animal ModelThe Animal Model of MI was created as described previously (Wang et al., 2018). Animals were anesthetized with an intraperitoneal treatment of pentobarbital sodium (50 mg/kg), and the chest was opened at the fourth intercostal space, and the heart was exposed. The rats in MI group were ligated the left anterior descending (LAD) coronary artery at 2-3 mm from its origin by a 6-0 polypropylene suture. The rats in the sham-operated group had no ligation and were considered as surgical controls. Some MI rats were treated with 50 g/kg/day ellagic acid by intragastric administration for 4 weeks. In short, the animals were divided into 3 groups as follows: (1) sham-operated control group (sham group, n = 6); (2) MI with vehicle group (MI group, n = 6); (3) MI+Ellagic acid group (50 mg/kg/day, n = 6).
Masson’s Trichrome StainingMasson’s Trichrome Staining assay was used to assess the cardiac fibrosis area. Four weeks after the MI surgery, the rats were anesthetized with pentobarbital sodium as described above and then were sacrificed. The left ventricles were separated and fixed in 4% paraformaldehyde. Then, tissues were embedded in paraffin and 5-6 μm tissue sections were prepared. Paraffin sections were dewaxed and washed with distilled water, and then dyed with hematoxylin for 2 min. Next, paraffin sections were rinsed with distilled water once again and dyed with ponceau staining solution for 15 min. Later, they were dyed with aniline blue (2.5 g aniline blue, 2 mL glacial acetic acid and 100 mL distilled water) for 5 min and washed with 1% glacial acetic acid solution for another 5 min. Finally, they were dehydrated by gradient alcohol and xylene, and mounted by resinene. The stained sections were used to conduct microscopic examination by a microscope (Olympus, Tokyo, Japan). The region stained green revealed cardiac fibrosis, and the red area indicated non-ECM myocardium. Images were acquired at 400 × magnification and analyzed using Image-Pro Plus 6.0 software (Media Cybernetics, Rockville, MD, USA). The cardiac fibrosis area was calculated as the fibrotic area stained green divided by the total area of the section (Wang et al., 2018; Pan et al., 2012).
Isolation and Culturing of CFsCFs were isolated from the left ventricles of the rats as described previously (Wang et al., 2018). In simple terms, the issue of left ventricles were minced and digested with 0.05% collagenase II (Sigma, St. Louis, MO, USA). The CFs were pre-plated for 1 hr on uncoated culture discs and rapidly adhered to the dishes. The adherent CFs were isolated after centrifugation and resuspension. Then, the CFs were cultured in Dulbecco’s modified Eagle’s medium (DMEM) containing 10% fetal calf serum (FCS, Gibco, Grand Island, NY, USA) and gentamycin (0.005%, Gibco) at 37°C and 5% CO2. The cells of passages 2-3 were used in the present study.
RNA extraction and real-time quantitative PCRTotal RNA was extracted with TRIzol Reagent (Sigma) and further treated with RNase-free DNase I (Promega, Madison, WI, USA) to remove genomic DNA contamination. A NanoDrop spectrophotometer and 1% agarose gels electrophoresis were used to determine the concentration and quality of RNA samples. The cDNA was obtained from 500 ng total RNA with the Iscript cDNA synthesis kit (Invitrogen, Carlsbad, CA, USA). Quantitative real-time PCR (qRT-PCR) was performed with an iCycler Real-Time PCR detection system (Biorad, Hercules, CA, USA) using iQ SYBR-Green supermix (Biorad). For all primer sets used (Table 1), temperature and dilution curves were assessed to check for linearity. The house-keeping gene CYP was used to normalize the relative genes expression of HDACs, collagen I/III, MMP-2 and MMP-9.
| Target Genes | Primers | Sequence (5’ to 3’) |
|---|---|---|
| Collagen I | Forward | TGGTCTTGGAGGAAACTTTGC |
| Reverse | CTGTGTCCCTTCATTCCGG | |
| Collagen III | Forward | ACCTGAAATTCTGCCACCCT |
| Reverse | GCCTTGAATTCTCCCTCATTG | |
| MMP-2 | Forward | GGAAGCATCAAATCGGACTG |
| Reverse | CACCCTCTTAAATCTGAAATCACC | |
| MMP-9 | Forward | AAGGATGGTCTACTGGCACA |
| Reverse | TTGCGTTTCCAAAGTAAGTG | |
| CYP | Forward | TATCTGCACTGCCAAGACTGAGTG |
| Reverse | CTTCTTGCTGGTCTTGCCATTCC |
Tissues or CFs were lysed with ice-cold RIPA lysis buffer containing a protease inhibitor (CWBIO, Beijing, China). Following a 10% SDS-PAGE gel separation, the protein was transferred to polyvinylidene fluoride membrane (PVDE). Then the PVDF were incubated with primary antibodies: anti-HDAC1 (1/1000), anti-HDAC2 (1:10000), anti-HDAC6 (1:3000), anti-Collagen I (1:2500), anti-Collagen III (1:5000), anti-MMP-2 (1:1000), anti-MMP-9 (1:500), or anti-β-actin (1:5000) at 4°C for 24 hr. Subsequently, the blots were incubated with secondary antibodies at room temperature for 1 hr. Specific bands were visualized using the ECL system on the Bio-Rad electrophoresis image analyzer (Biorad) and quantified using ImageJ software. All antibodies used in the present study were purchased from Abcam Company. The protein of β-actin was considered as a reference to normalize the density of other protein bands in this study.
Proliferation AssayThe CFs were cultured in 96-well plates. After serum-starvation for 24 hr, the CFs were treated with or without Ang II, Ang II plus Ellagic acid. Then, Cell Counting Kit-8 (Sigma) was used to analyze the proliferation of CFs according to the manufacturer’s instructions. The optical density was detected at an absorbance of 450 nm by a microplate reader (Biorad).
Migration AssayThe CFs were incubated in the upper Transwell chambers in serum-free DMEM. DMEM (containing serum) with or without Ang II, Ang II plus Ellagic acid, was added to the lower Transwell chamber as a chemotactic stimulus, and the number of migrated CFs was counted at 8 hr after treatment.
Cell transfectionThe CFs were cultured in 96-well plates with a concentration of 2 × 104 cells/well for 24 hr. pcDNA-HDAC1 and its negative control pcDNA were transfected into CFs using Lipofectamine 2000 (Invitrogen), respectively.
Statistical analysisThe data were analyzed using SPSS 17.0. Independent T-test was used to compare the difference between the two groups. All data are shown as mean ± SD. P < 0.05 indicates significant differences of data.
Firstly, we examined the genes expression of HDACs and fibrosis-related factors in the heart tissues of MI rats. Compared to sham rats, the protein levels of HDAC1, HDAC2 and HDAC6 (Fig. 1A) and the mRNA levels of collagen I, collagen III, MMP-2 and MMP-9 (Fig. 1B, P < 0.05) were significantly up-regulated in MI rats. In parallel, the Masson result also showed that the area of cardiac fibrosis in MI rats was significantly increased compared to sham rats (Fig. 1C, P < 0.05). However, ellagic acid decreased the protein level of HDAC1 (Fig. 1A) and the mRNA expression of fibrosis-related genes (Fig. 1B, P < 0.05), and reduced the area of cardiac fibrosis (Fig. 1C, P < 0.05) in MI rats.
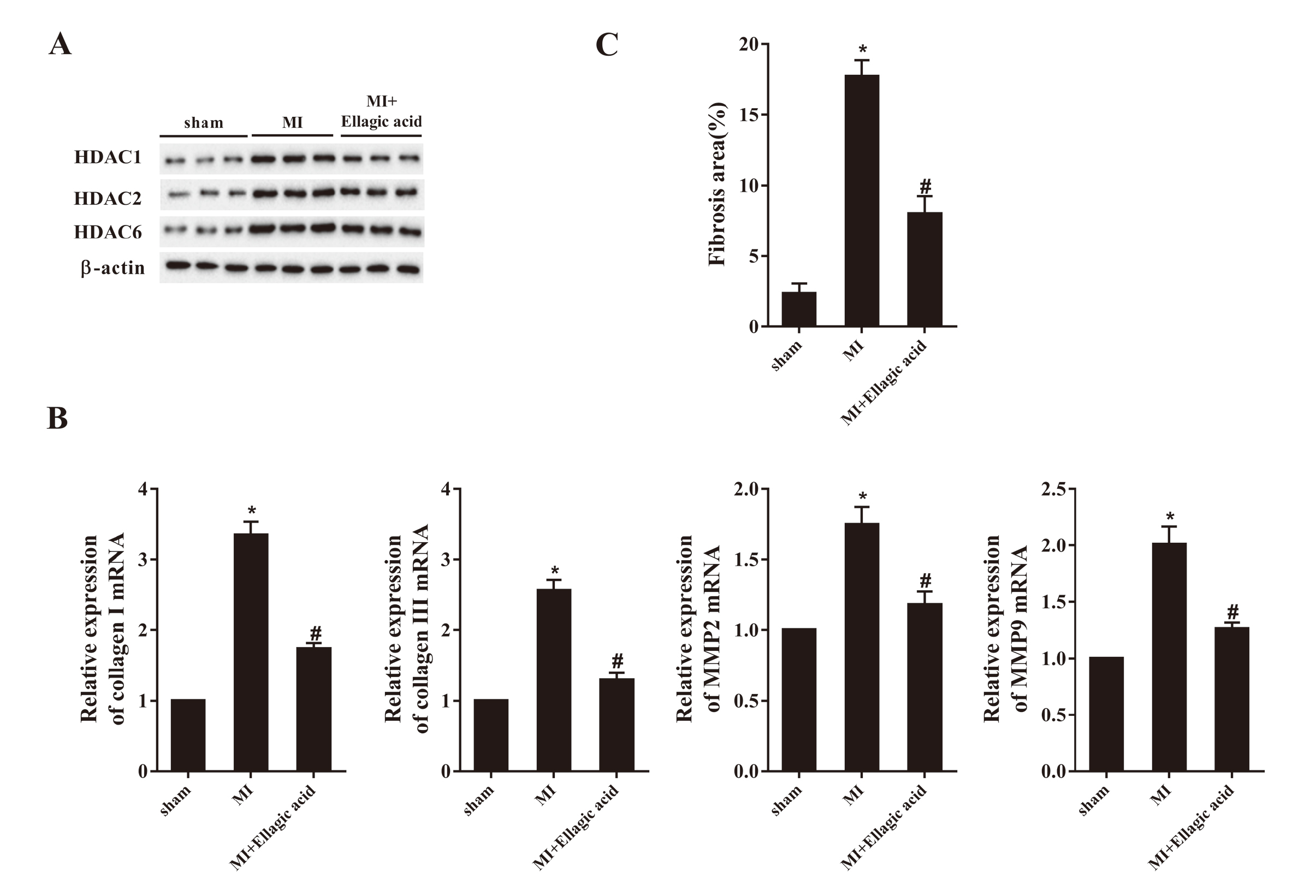
Ellagic acid down-regulated the gene expression of HDAC1, collagen I/III and MMP2/9, and attenuated cardiac fibrosis in MI rats. The rats were divided into 3 groups (n = 6): sham group, MI group and MI+Ellagic acid group. A: Western blot images showing the effects of Ellagic acid on the expression of HDACs. β-actin was used as a reference. B: the mRNA expression profiles of collagen I/III and MMP2/9. C: Quantitative analysis of cardiac fibrosis evaluated by Masson’s trichrome staining. *P < 0.05 vs. sham; #P < 0.05 vs. MI+Ellagic acid.
Further, CFs were stimulated by 100 nM Ang II for 24 hr and the impacts of ellagic acid on HDAC expression were assessed. The WB results revealed that Ang II stimulation significantly increased the protein expression of HDAC1 compared to control, while 60 μmol/L and 120 μmol/L ellagic acid reversed the effects (Fig. 2A & 2B, P < 0.05).
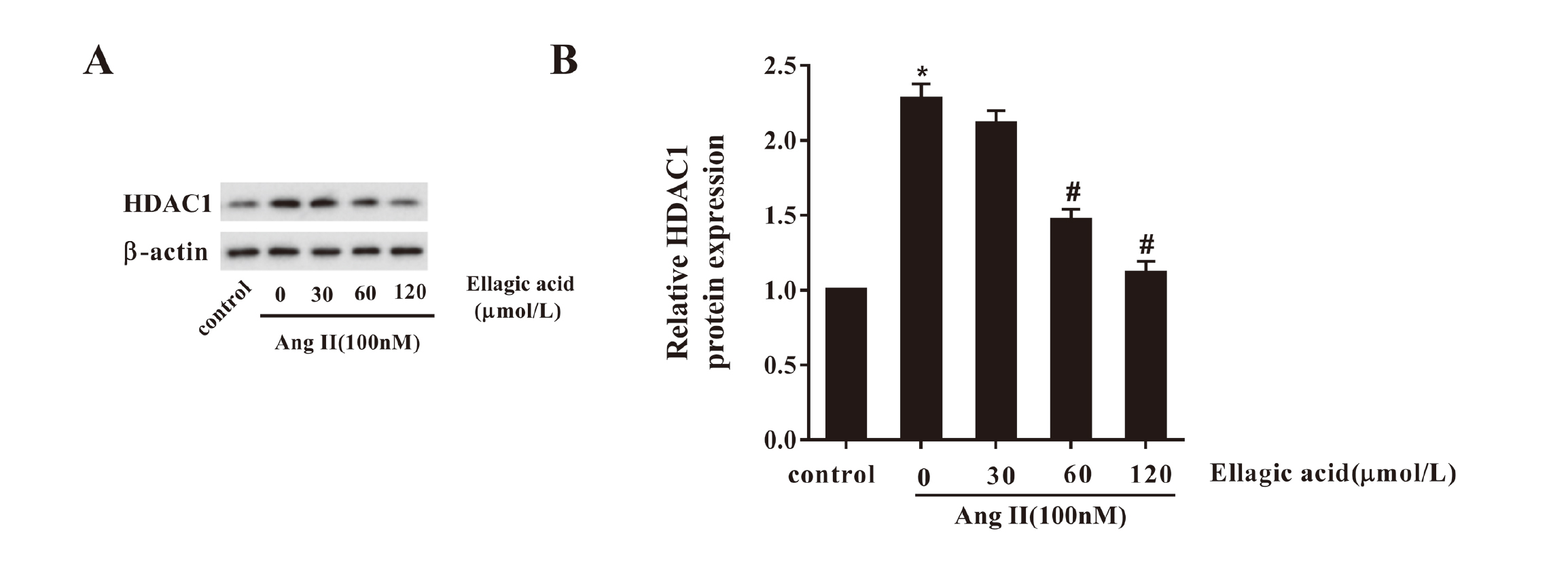
Ellagic acid repressed the protein expression of HDAC1 in CFs. Five different treatments were conducted to CFs as follows: control, Ang II, Ang II +30 μmol/L Ellagic acid, Ang II +60 μmol/L Ellagic acid and Ang II +120 μmol/L Ellagic acid. Therein, the concentration of Ang II was 100 nM, and the stimulation lasted 24 hr. A: Western blot images showing the effects of Ellagic acid on the expression of HDAC1. B: Quantitative analysis of the HDAC1 protein expression measured by western blot. *P < 0.05 vs. sham; #P < 0.05 vs. Ang II.
We detected the variation of gene expression of fibrosis-related factors in CFs following the stimulation of 100 nM Ang II for 24 hr. Consistent with the change of the HDAC1 expression, stimulation of Ang II significantly induced the protein expression of fibrosis-related factors collagen I, collagen III, MMP-2 and MMP-9 compared to control, while 60 μmol/L and 120 μmol/L ellagic acid reversed the effects (Fig. 3A & 3B, P < 0.05).
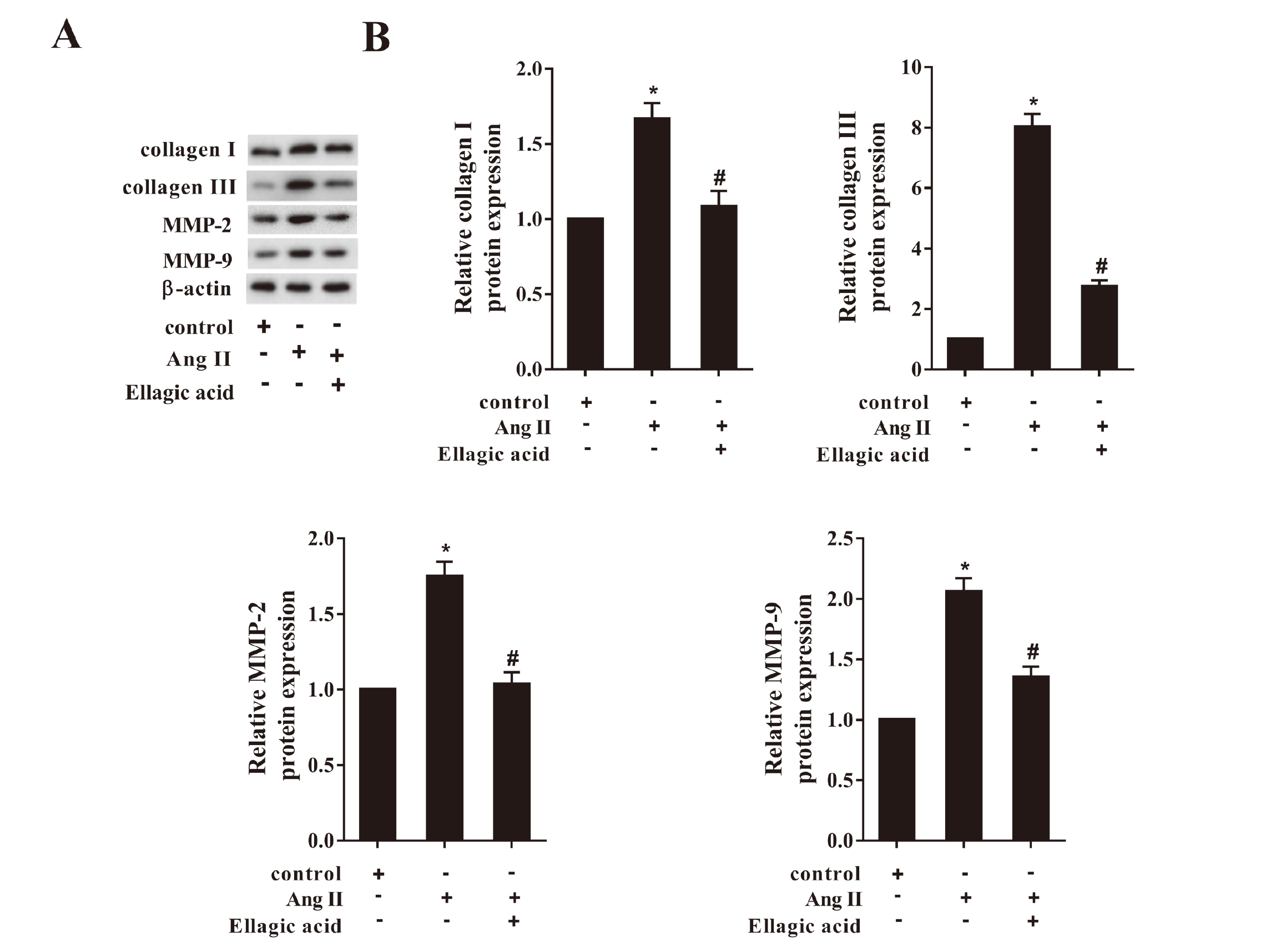
Ellagic acid suppressed the expression of the fibrosis-related factors in CFs. The CFs were divided into 3 groups: control, Ang II and Ang II+ Ellagic acid. The concentration of Ang II was 100 nM, and the stimulation lasted 24 hr. A: Western blot images showing the effects of Ellagic acid on the expression of the fibrosis-related factors. B: Quantitative analysis of collagen I/III, MMP-2 and MMP-9 protein expressions detected by western blot. *P < 0.05 vs. control; #P < 0.05 vs. Ang II.
We measured the effects of ellagic acid on proliferation and migration of CFs after Ang II stimulation by CCK8 and Transwell assays. The OD value revealed that Ang II stimulation significantly accelerate the proliferation of CFs, while ellagic acid abolished this trend (Fig. 4A, P < 0.05). Similarly, the Transwell results showed that Ang II remarkably promoted the migration of CFs, while ellagic acid eliminated this effect (Fig. 4B, P < 0.05).
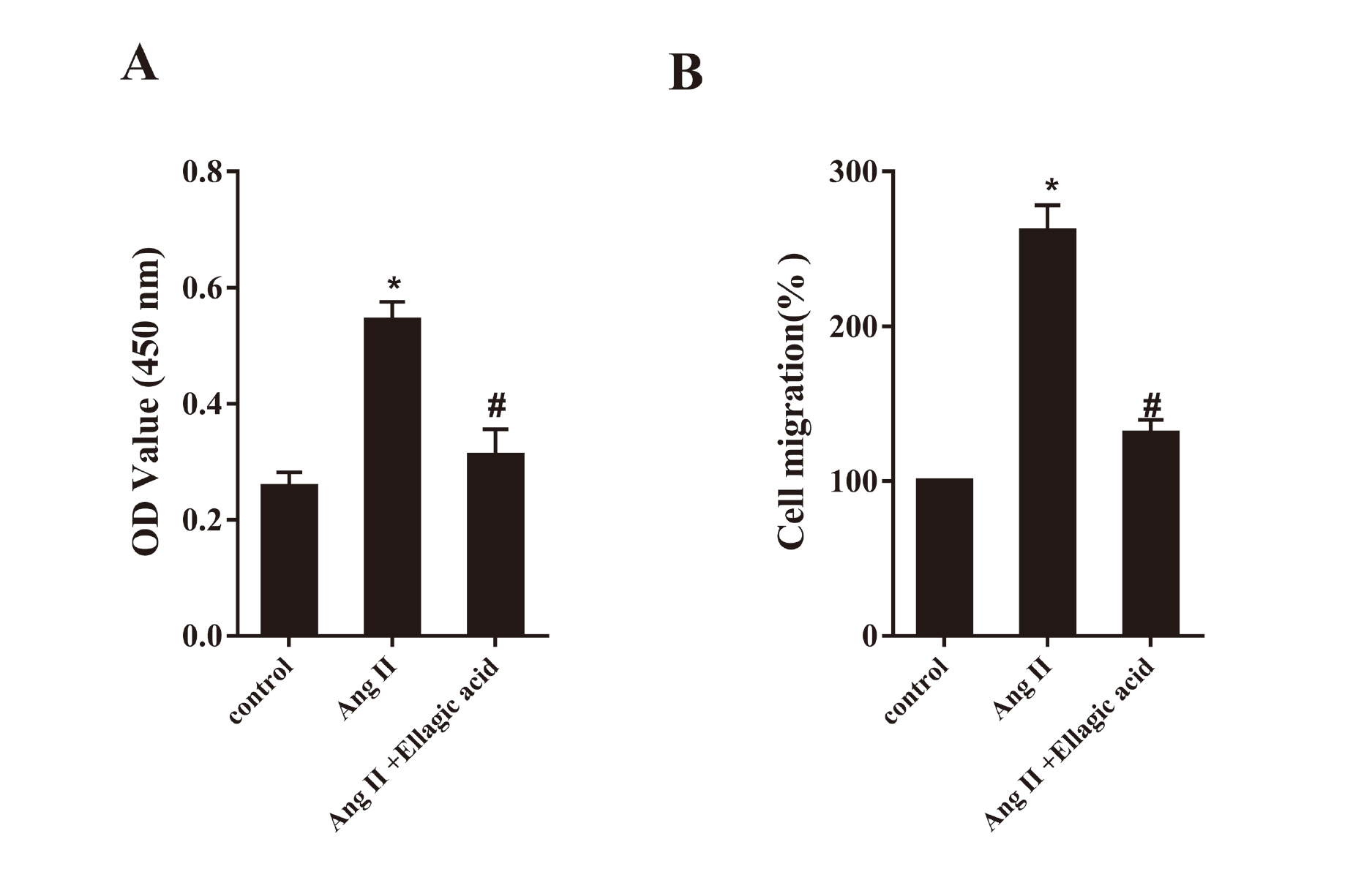
Ellagic acid inhibited Ang II-stimulated proliferation (A) and migration (B) of CFs. The CFs were divided into 3 groups: control, Ang II and Ang II+ Ellagic acid. The concentration of Ang II was 100 nM, and the stimulation lasted 24 hr. *P < 0.05 vs. control; #P < 0.05 vs. Ang II.
CFs were transfected with pcDNA-HDAC1 or negative control pcDNA. Then we examined the protein levels of fibrosis-related factors, and the results were shown in the Fig. 5A and 5B. Ellagic acid remarkably suppressed the protein expression of collagen I, collagen III, MMP-2 and MMP-9, while HDAC over-expression canceled this impact.
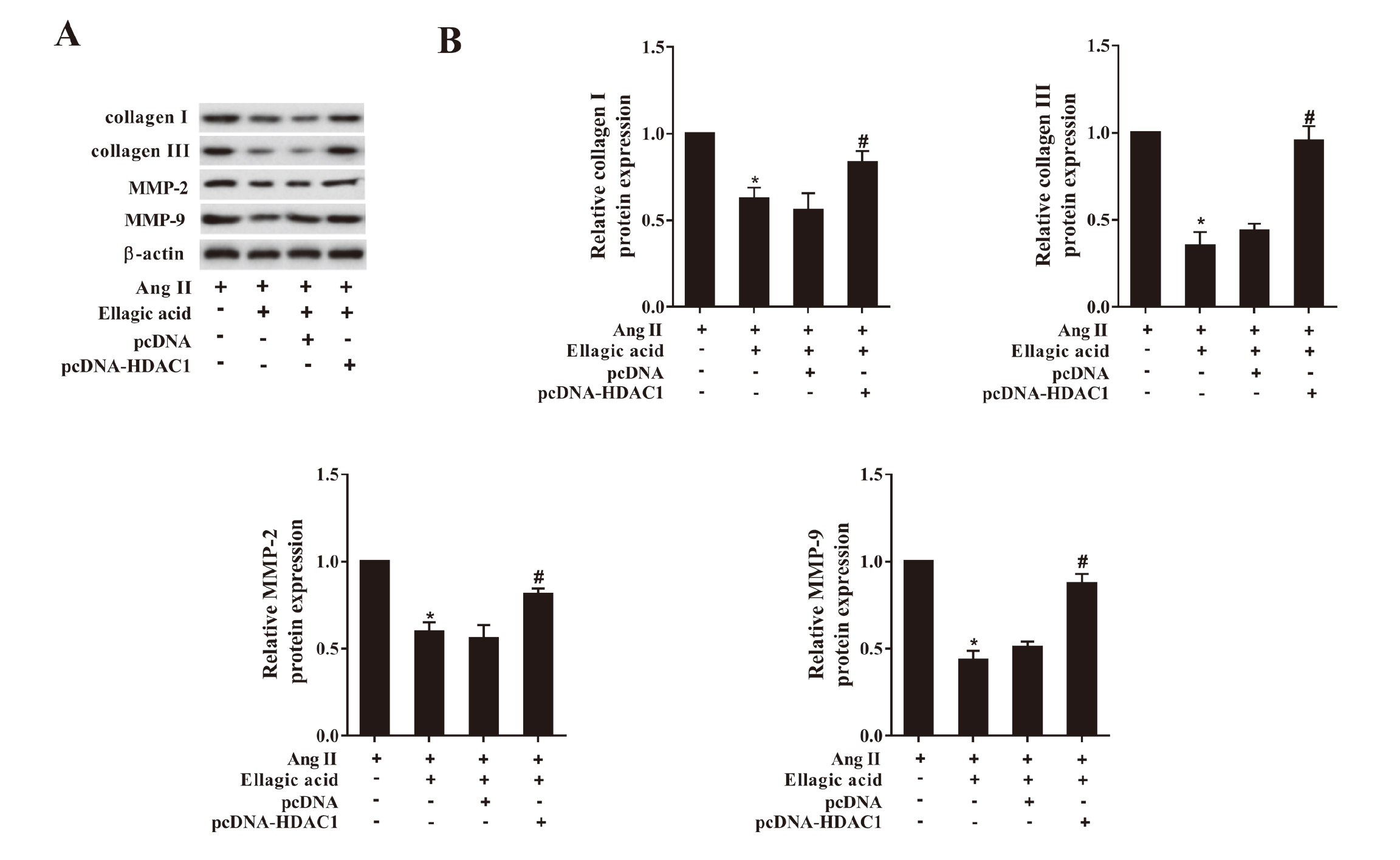
Ellagic acid regulated the protein expression of fibrosis-related factors through HDAC1. The CFs were treated under the following conditions: Ang II, Ang II +Ellagic acid, Ang II +Ellagic acid+ pcDNA and Ang II +Ellagic acid+ pcDNA-HDAC1. The dose of Ang II was 100 nM, and the CFs were stimulated for 24 hr. The concentration of Ellagic acid was 60 μmol/L. A: Western blot images showing the effects of Ellagic acid on the expression of the fibrosis-related factors. B: Quantitative analysis of collagen I/III, MMP-2 and MMP-9 protein expressions detected by western blot. *P < 0.05 vs. Ang II; #P < 0.05 vs. Ang II +Ellagic acid+ pcDNA-HDAC1.
As described above, CFs were transfected with pcDNA-HDAC1 or negative control pcDNA. Similarly, the results exhibited that ellagic acid inhibited the proliferation (Fig. 6A, P < 0.05) and migration (Fig. 6B, P < 0.05) of CFs, and those effects were abrogated by over-expressing HDAC1.
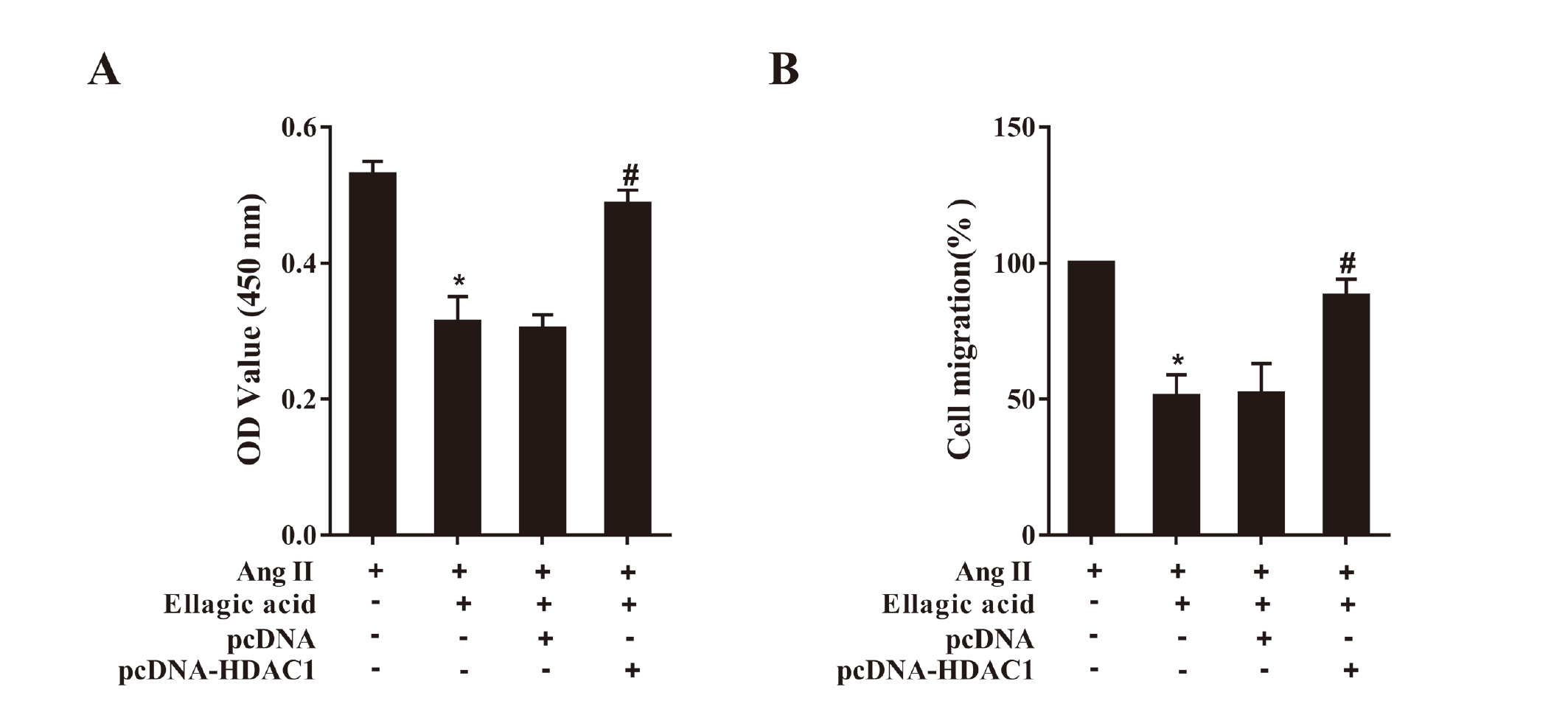
Ellagic acid inhibited the proliferation (A) and migration (B) of CFs through HDAC1. The CFs were treated under the following conditions: Ang II, Ang II +Ellagic acid, Ang II +Ellagic acid+ pcDNA and Ang II +Ellagic acid+ pcDNA-HDAC1. The dose of Ang II was 100 nM, and the CFs were stimulated for 24 hr. The concentration of Ellagic acid was 60 μmol/L. Ang II; #P < 0.05 vs. Ang II +Ellagic; #P < 0.05 vs. acid+ pcDNA-HDAC1.
Collagens, the secreta of CFs, are the major components of ECM; and the matrix metalloproteinase (MMPs) are key enzymes to involve in the degradation and synthesis of ECM (Yabluchanskiy et al., 2013). In the current study, we found that the mRNA levels of collagen I, collagen III, MMP-2 and MMP-9 in MI rats and the protein expression of these genes in Ang II-stimulated CFs were both significantly elevated. The present results were in accordance with a previous study conducted on rats (Wang et al., 2018). In addition, the area of cardiac fibrosis in MI rats was larger than in sham rats. That corroborated that MI promoted the cardiac fibrosis in rats.
Recent evidence has revealed that ellagic acid could be a novel drug in therapy of MI. Previous research showed that ellagic acid decreased infarct size in MI rats (Mari Kannan and Darlin Quine, 2012). Also, oral pretreatment with ellagic acid restored pathological arrhythmias in MI rats (Kannan and Quine, 2013). Further, our previous study indicated that ellagic acid improved ventricular remodeling after MI by regulating the miR-140-3p/MKK6 pathway (Wei et al., 2017). Similarly, we found that ellagic acid decreased expression of fibrosis-related factor in MI rats, reduced the area of cardiac fibrosis and inhibited the proliferation and migration of CFs. Our results were consistent with those previous studies. That suggested that ellagic acid could relieve cardiac fibrosis caused by MI.
Accumulating evidence shows that HDACs play a critical role in tumor growth and cardiovascular diseases, especially class I HDACs (Montgomery et al., 2007). In the present study, we observed that the protein expression of HDACs (HDAC1, HDAC2 and HDAC6) in the MI rats and HDAC1 in Ang-II stimulated CFs were both significantly up-regulated. Similarly, a previous study shown that the protein levels of HDAC1 and HDAC2 were both more abundant in MI rats than in sham rats (Nural-Guvener et al., 2014). Another study indicated that the expression of HDAC6 was induced in cardiac fibrotic tissue and activated CFs (Pan et al., 2012). Those parallel results suggested the expression of HDACs could promote the cardiac fibrosis.
Due the impacts of HDACs, several studies focused on the function of HDAC inhibitors in MI and other diseases. According to the results of those studies, HDACs inhibitors could cause cell cycle arrest (Lee et al., 2007) and reduce cardiac hypertrophy in pathological conditions (Cardinale et al., 2010). Furthermore, HDACs inhibitors enhanced cardiac contractility and attenuated structural remodeling in heart failure (Kao et al., 2013).
Interestingly, we found that ellagic acid inhibited expression of HDAC1 in MI rats and Ang-II-stimulated CFs in the present study. That might suggest ellagic acid could negatively regulate the expression of HDAC1. Compared to pcDNA, the suppressed effects of ellagic acid on proteins expression of fibrosis-related genes were remarkably reversed when HDAC1 over-expressed. The results of proliferation and migration showed the same variations in the Ang-II stimulated CFs. An experiment implemented on rats discovered that the silence of HDAC6 inhibited expression of TGF-xp and reduced the proliferation of CFs (Tao et al., 2016). Those results might reveal that ellagic acid could inhibit proliferation and migration of CFs and attenuate cardiac fibrosis by down-regulating expression of HDACs.
In summary, ellagic acid could inhibit the expressions of fibrosis related-factors in MI rats and Ang-II stimulated CFs, reduce the area of cardiac fibrosis in MI rats, and decrease the proliferation and migration of CFs through suppressing expression of HDAC1. The present study suggested the effective anti-fibrosis effect of ellagic acid in MI and also provided a worthwhile therapeutic target of MI in the future.
This study was supported by grant from the Medical Health Science and Technology Project of Zhejiang Provincial Health Commission (NO. 2018KY526).
Conflict of interestThe authors declare that there is no conflict of interest.