2021 Volume 10 Issue 1 Pages A0096
2021 Volume 10 Issue 1 Pages A0096
The gas-phase adsorption of N2 on protonated serine (Ser, C3H7NO3), threonine (Thr, C4H9NO3), glycine (Gly, C2H5NO2), and 2-aminoethanol (C2H7NO) was investigated using a tandem mass spectrometer equipped with an electrospray ionization source and a cold ion trap. N2 molecules were adsorbed on the free X–H (X=O and N) groups of protonated molecules. Gas-phase N2 adsorption-mass spectrometry detected the presence of free X–H groups in the molecular structures, and was applied to the structural elucidation of small molecules. When the 93 structures with an elemental composition of C3H7NO3 were filtered using the gas-phase N2 adsorption-mass spectrometry results for Ser, the number of possible molecular structures was reduced to 8 via the quantification of the X–H groups. Restricting and minimizing the number of possible candidates were effective steps in the structural elucidation process. Gas-phase N2 adsorption-mass spectrometry combined with mass spectrometry-based techniques has the potential for being useful for elucidating the molecular structures of a variety of molecules.
Molecular structures are typically identified using nuclear magnetic resonance spectroscopy, X-ray crystallography, optical spectroscopy, and chromatography. Mass spectrometry is routinely used for structural elucidation owing to its high sensitivity, selectivity, and suitability for analyzing mixtures. The elemental compositions of unknown molecules are assigned using high-resolution and accurate mass spectra collected for complex mixtures.1) The number of potential elemental compositions, which increases largely with mass, can be reduced by comparing the experimental and theoretical isotope patterns in mass spectra.2)
When the elemental composition of an analyte is elucidated using mass spectrometry, the molecular structure is examined utilizing product ion profiles that are obtained via tandem mass spectrometry. The relationships between molecular structure and fragmentation have been studied using a variety of dissociation techniques.3,4) Ion mobility and hydrogen/deuterium exchange have been used to analyze molecular structures.5,6) Nuclear magnetic resonance spectroscopy of gas-phase ions using a magnetic resonance acceleration technique has been developed.7) Mass spectral libraries and electronic databases are typically surveyed for the structural elucidation of unknown molecules in complex mixtures, because the number of potential molecular structures with any elemental composition increases exponentially with mass.8–10)
The molecular adsorption of gas-phase ions provides information on the structure of gas-phase ions. N2 adsorption on small metal clusters of cations depends on the structure of the cations.11,12) Molecular adsorption experiments on gas-phase carbohydrate ions with 2–5 hydroxy (O–H) groups revealed that the O–H groups of carbohydrates could be quantified using gas-phase N2 adsorption-mass spectrometry.13)
In this study, we investigated the adsorption of gas-phase N2 on protonated molecules using a tandem mass spectrometer equipped with an electrospray ionization source and temperature-controlled ion trap. Gas-phase N2 adsorption-mass spectrometry was used for structural elucidation in addition to assigning elemental composition and structural elucidation using fragmentation and mass spectral library surveys with serine (Ser, C3H7NO3), threonine (Thr, C4H9NO3), glycine (Gly, C2H5NO2), and 2-aminoethanol (C2H7NO) being used as sample molecules.
Gas-phase N2 adsorption was detected using a home-built tandem mass spectrometer equipped with an electrospray ionization source and a temperature-controlled 22-pole ion trap (8–350 K).14,15) Protonated molecules were generated via the electrospray ionization of solutions containing 0.5 mM of an analyte molecule in a mixture of water and methanol with 1% acetic acid. Analyte molecules were obtained from Nacalai Tesque. The ionization source was operated at a sample flow rate of 3 μL/min, a sheath gas flow rate of 3 L/min, and an applied voltage of 1.5 kV. The protonated molecules were transferred to the gas phase through a metal capillary (340 K) and stored in an octopole ion guide. The gas-phase ions were pulsed into a quadrupole mass filter and a temperature-controlled 22-pole ion trap. The mass-selected ions were thermalized via multiple collisions with He containing 30% N2 as the buffer gas for 50 ms in the ion trap at 60 K, and the N2 molecules were adsorbed on the mass-selected ions in the ion trap. At lower temperatures, the N2 molecules condensed on the ion trap electrodes, while at room temperature, the N2 molecules were not adsorbed on the ions. The temperature of the ion trap was controlled using a cryogenic refrigerator (CH-204SB, Sumitomo, Japan) and a heater cartridge (HTR-50, Lake Shore, USA), and measured using two silicon diode temperature sensors (DT-670B-CU, Lake Shore, USA). The exact partial pressure and temperature of the N2 molecules in the ion trap were not determined in this study, and the ambient pressure in the chamber containing the ion trap was approximately 2×10−4 Pa. The maximum numbers of N2 molecules that were adsorbed did not depend on the chamber pressure, but the efficiency of N2 adsorption was dependent on this. The N2-adsorbed cluster ions were extracted from the ion trap and orthogonally accelerated to 2.8 keV using two-stage pulsed electric fields. The ions were mass-analyzed using a reflectron time-of-flight (TOF) spectrometer and counted using dual microchannel plates (F4655, Hamamatsu Photonics, Japan) and a digital storage oscilloscope (104MXi, LeCroy, Japan). The desorption of N2 from the weakly bound clusters in the field-free region of the reflectron TOF mass spectrometer was also observed in the TOF mass spectra, in addition to the N2-adsorbed cluster ions that were formed in the ion trap.15) The repetition rate of the experimental cycle was 10 Hz.
Figure 1a illustrates the mass spectrum of H+Ser(N2)n, which formed via the adsorption of N2 on mass-selected H+Ser at 60 K. Here, n denotes the number of N2 molecules that are adsorbed on the H+Ser in the ion trap, which, according to our experimental results, was n≤5. This indicates that the first solvation shell of H+Ser consisted of a maximum of five molecules. Stable magic number clusters formed via gas-phase N2 adsorption on mass-selected ions were observed for carbohydrates and tryptophan (Trp).13,14) It was reported that N2 molecules are adsorbed on the H atoms of the O–H groups of carbohydrates, which allowed for the O–H groups of carbohydrates to be quantified using gas-phase N2 adsorption data.13) H+Trp(N2)5 was a stable magic number cluster in which five N2 molecules were adsorbed on the H atoms of the NH3+ group, the indole ring, and the carboxyl (COOH) group.14) H+Trp(H2)n with n=1–5 were reported to be formed in messenger-tagging vibrational predissociation spectroscopic measurements of H+Trp using a cryogenic ion trap.16) Diatomic molecules were primarily bound by charge-induced dipole interactions. The optimized structure of H+Ser(N2)5 at the B3LYP/6-31++G(d,p) level using the Gaussian 09 package17) was obtained utilizing previously reported data14,16,18) (Fig. 1b), and the results suggested that five N2 molecules were adsorbed on the NH3+, O–H, and COOH groups of the H+Ser. The relative ion intensity of H+Ser(N2)5 was lower than that of H+Ser(N2)4, as illustrated in Fig. 1a. The ion intensity distributions of the N2-adsorbed cluster ions indicated that the gas-phase ions had multiple structures.13) The lower relative ion intensity of the n=5 cluster was attributed to the intramolecular hydrogen bond between the NH3+ group and the oxygen atom of COOH group as illustrated in Fig. 1b. In the case of Thr, like Ser, contains an O–H group, the maximum number of adsorbed N2 molecules was n=5 as illustrated in Fig. 2, indicating that the adsorption sites of H+Thr were the NH3+, O–H, and COOH groups as in the case of H+Ser. To reveal the effects of molecular adsorption on the geometric and electronic structures of gas-phase ions, photodissociation spectroscopy and theoretical calculations are required.


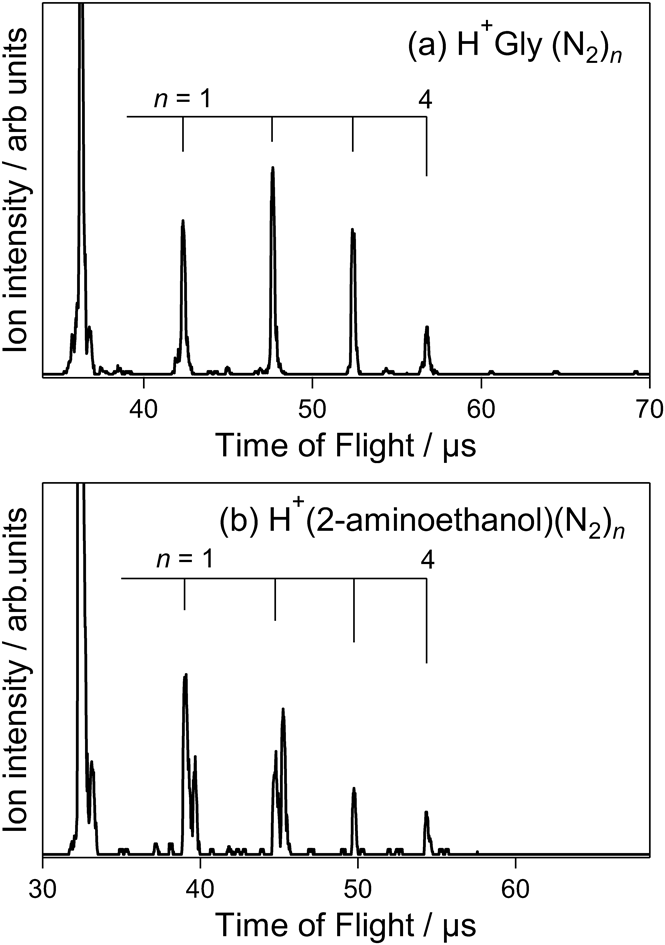
To confirm the relationship between N2 adsorption and molecular structure, gas-phase N2 adsorption-mass spectrometry of H+Gly and H+(2-aminoethanol) was performed. Gly and 2-aminoethanol lack the O–H and COOH groups of Ser, respectively. Figure 3a illustrates the mass spectrum of H+Gly(N2)n, which forms via the adsorption of N2 on mass-selected H+Gly at 60 K. The maximum number of adsorbed N2 molecules was n=4, and an n=5 cluster was not observed in the spectrum. The number of adsorption sites for H+Gly was one less than that of H+Ser and H+Thr. This indicates that four N2 molecules were adsorbed on the NH3+ and COOH groups of H+Gly. The maximum number of adsorbed N2 molecules for H+(2-aminoethanol)(N2)n was also n=4 (Fig. 3b), indicating that four N2 molecules were adsorbed on the NH3+ and O–H groups of H+(2-aminoethanol). The above findings indicate that gas-phase N2 adsorption-mass spectrometry can be used to detect the presence of free X–H (X=O and N) groups in molecular structures.
Based on the aforementioned results, we proposed a method for structural elucidation using gas-phase N2 adsorption-mass spectrometry that can be used prior to fragmentation and in surveying mass spectral libraries. Figure 4 illustrates the strategy used for the structural characterization of Ser (C3H7NO3, calculated exact mass 105.0426). The CAS registry revealed 93 possible structures with an emperical formula of C3H7NO3. When conventional methods are used for characterizing molecular structures, the product ion spectrum of the protonated molecule is compared with the product ion spectra in a mass spectral library, and the structures that match the elemental composition and fragmentations are screened.8-10) Restricting and minimizing the number of possible candidates are effective steps of the analysis process.
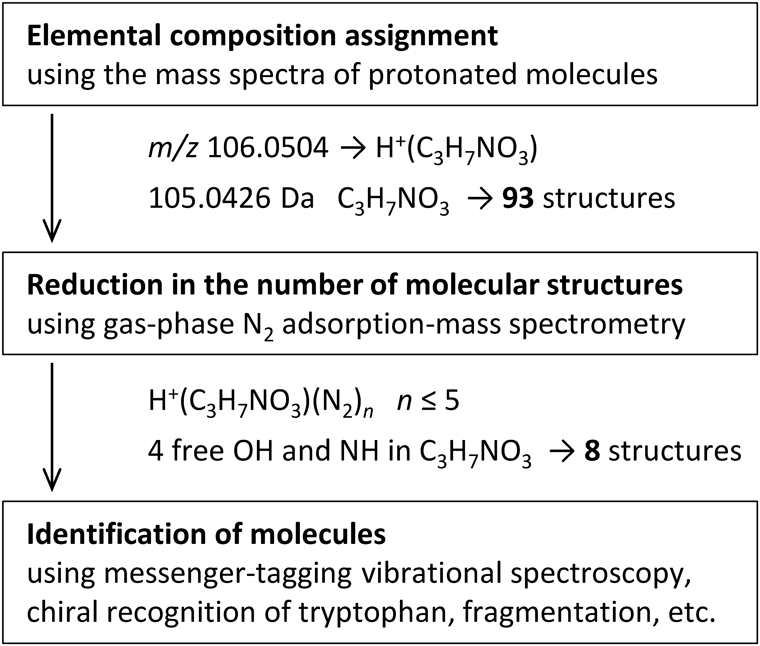
Gas-phase N2 adsorption-mass spectrometry can reduce the number of possible molecular structures with the same elemental composition via the quantification of the X–H (X=O and N) groups of the protonated molecule. For the C3H7NO3 elemental composition, the maximum number of N2 molecules adsorbed on the protonated molecule was n=5, which indicated that C3H7NO3 contained four X–H groups. The number of adsorption sites of C3H7NO3 was derived by subtracting of 1 from the maximum number n=5 of H+(C3H7NO3), because one N2 molecule adsorbed the proton. When the 93 structures associated with the elemental composition of C3H7NO3 were filtered using the number of adsorption sites for C3H7NO3, the number of possible molecular structures was reduced to eight (Fig. 5a). In the case of Gly (C2H5NO2, calculated exact mass 75.0320), the 46 possible structures for C2H5NO2 were reduced to 5 (Fig. 5b) via the quantification of the adsorption sites. This method decreased to 8.6 and 10.8% of the number of possible candidates obtained from the conventional methods for C3H7NO3 and C2H5NO2, respectively.

The gas-phase adsorption of N2 on protonated molecules generated via electrospray ionization was investigated using a tandem mass spectrometer equipped with a temperature-controlled ion trap. This method allowed the number of possible molecular structures with the same elemental composition to be reduced by tenth via the quantification of the X–H (X=O and N) groups. Restricting the number of possible candidates is an effective step in the structural elucidation process.
The weakly bound clusters of analyte ions and N2 formed in cold ion traps, such as H+(C3H7NO3)(N2), can be used for the messenger-tagging vibrational spectroscopy analysis of gas-phase analyte ions.19) The optical properties of Trp hydrogen-bonded with analyte ions in the gas phase can be used to identify and quantify the isomers and enantiomers of analyte molecules.20,21) An ion mobility spectrometer with radio-frequency electric fields has been developed for high-resolution and high-transmission efficiency analyses of small molecules.22) The findings reported herein indicate that gas-phase N2 adsorption-mass spectrometry combined with mass spectrometry-based techniques can be useful for elucidating molecular structures.
Mass Spectrom (Tokyo) 2021; 10(1): A0096