2017 Volume 6 Issue 3 Pages S0069
2017 Volume 6 Issue 3 Pages S0069
We compared two ionization methods, matrix assisted laser desorption/ionization (MALDI) and nanoparticle assisted laser desorption/ionization (Nano-PALDI) mass spectrometry (MS), for the analysis of amino acids derivatized with Py-Tag™ that consists pyrylium-based compound. Py-Tag is a useful stable derivatization reagent due to wide mass differences (using 13C as the sole stable labelling isotope). For Py-Tag labelled lysine, sensitive signals that showed less noise with a ten times higher sensitivity, showed a wider mass difference by Nano-PALDI MS compared to MALDI MS.
Nanoparticle assisted laser desorption/ionization mass spectrometry (Nano-PALDI MS) is a simple ionization method that eliminates, not only uneven distribution and different sizes of crystals, and the background peaks in the low mass range caused by organic matrices. In previous reports, various analytes such as drugs, lipids, peptides, synthetic polymers and proteins could be ionized and imaged by Nano-PALDI MS.1–5) The nanoparticles used in this study were synthesized by mixing aqueous solutions of 3d transition metal chlorides (MCl2·nH2O) and 3-aminopropyltriethoxysilane in a single step, resulting in particles with a metal oxide core coated with a functionalized silicate sheet and are able to ionize analytes. The nano-PALDI method has problems associated with sensitivity although it is a suitable method for ionizing and detecting low molecular weight compounds.
In the matrix assisted laser desorption/ionization (MALDI) method, chemical matrices are used to enhance the ionization of the high molecular weight analytes. Ion suppression can occur from matrix related ions in the low mass region (Fig. 1). We attempted to improve sensitivity in the low molecular region by LDI method, although we are able to measure the target compound by selecting an ionization method according to molecular weight.
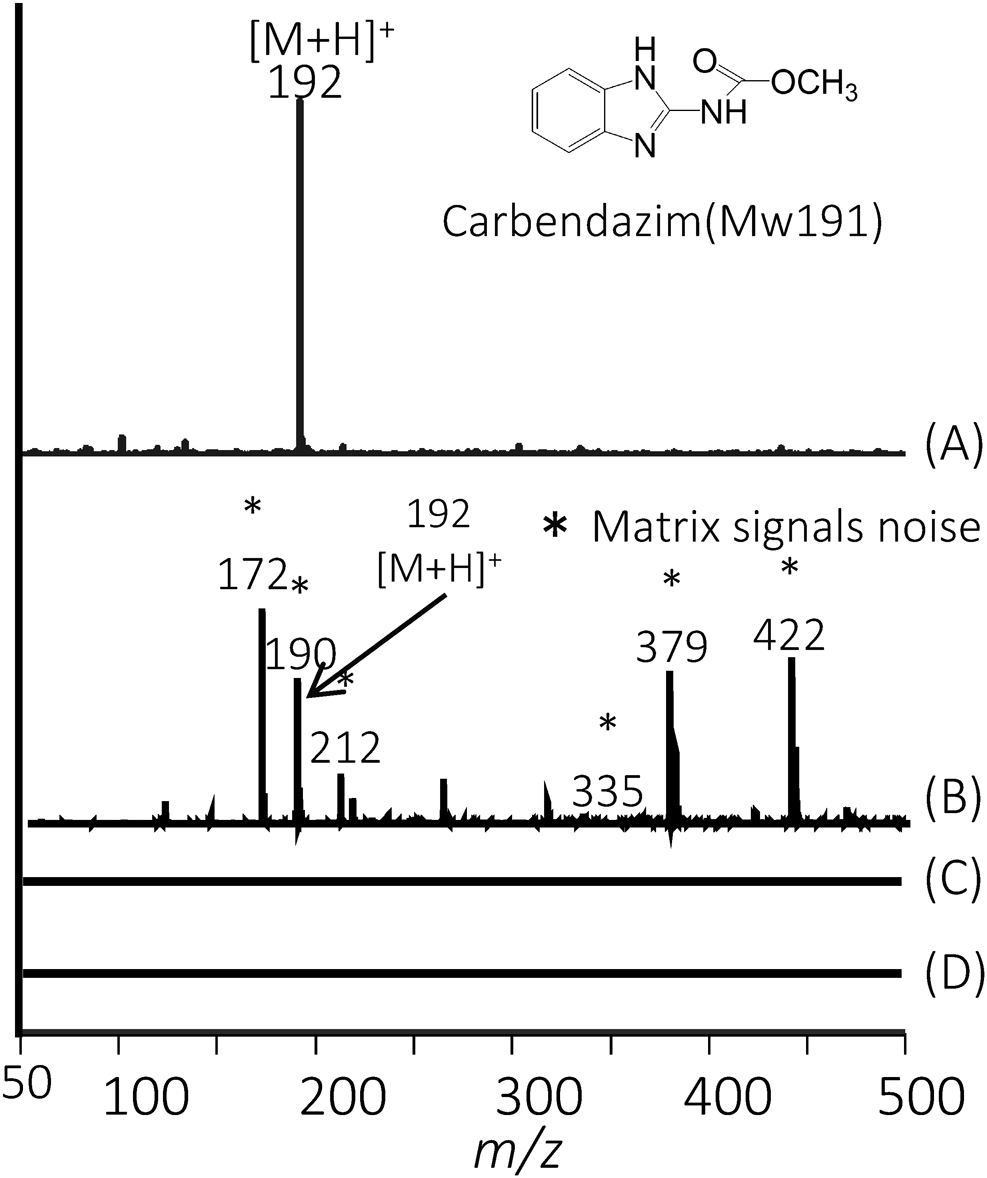
To enhance sensitivity, stable isotope-labelling reagents were developed to create mass differences for easy comparison of expression levels.6,7) As a new isotopic tagging reagent, we used Py-Tag**, which basically has a pyrylium structure and can specifically react with primary amino groups (e.g., the amino group of a lysine residue). Py-Tag increases the number of identifications due to its high labelling efficiency, wide mass difference, and chemical stability (Fig. 2A). Large mass differences can be created using Py-Tag using only 13C as a stable isotope without the need for other elements such as 15N or 18O.
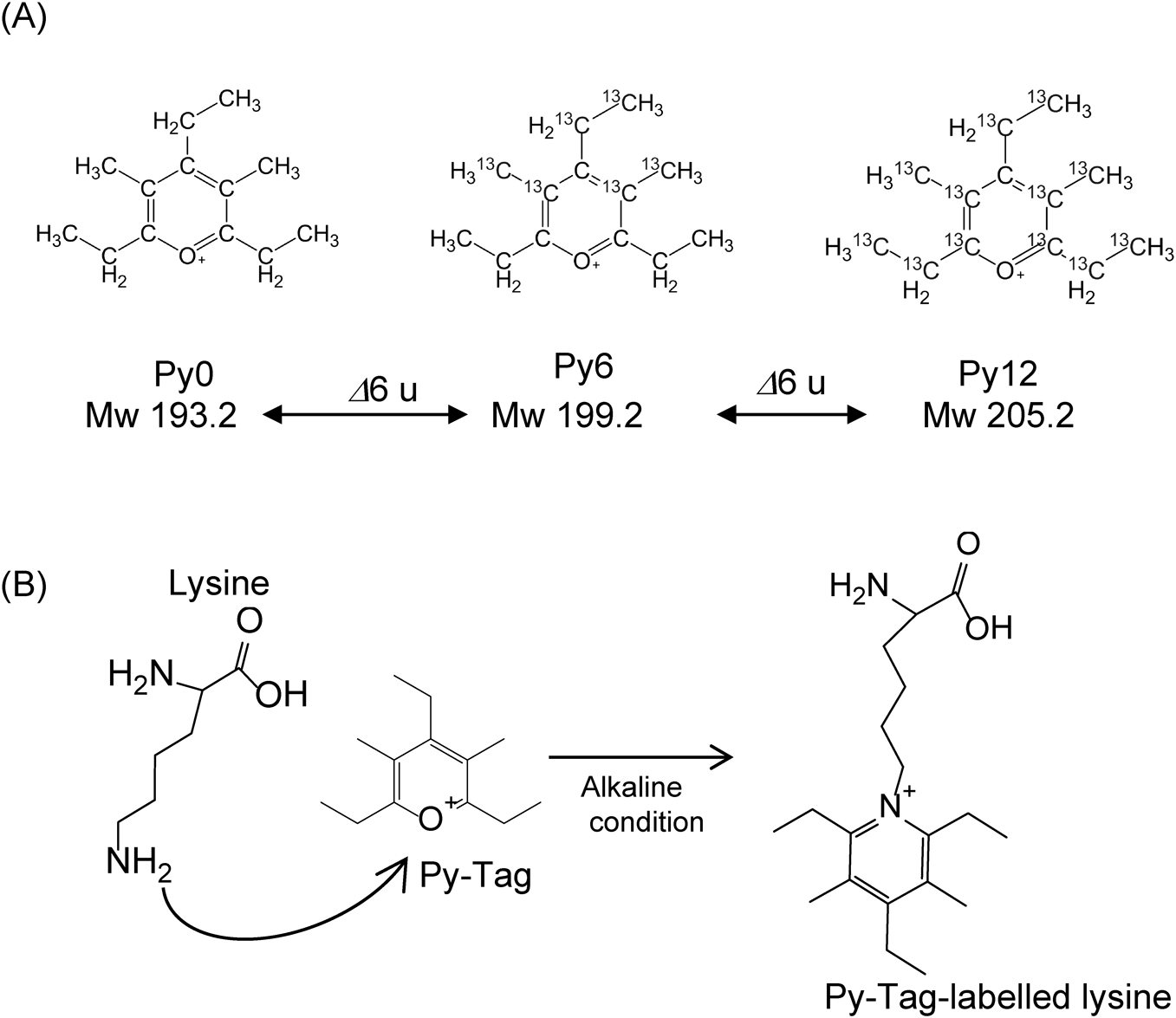
In this research, we attempted to find a better ionization method for increasing the sensitivity of detection of an amino acid compared with MALDI- or Nano-PALDI-coupled with Py-Tag labelling mass spectrometry (Fig. 2B).
The following aqueous solutions were prepared: lysine: 29.4 mg/mL, glycine: 15 mg/mL (Wako Pure Chemical Industries, Ltd., Japan). A 2.5 μL aliquot of mixed amino acid solution was mixed with 7.5 μL of reaction borate buffer (pH 9.5) in a sealed 0.2 mL PCR test tube. A 2.5 μL aliquot of Py0, 6 or 12 (Taiyo Nippon Sanso Co., Japan) (200 mM) was added, and the mixture was then heated at 55°C for 4 h. A 0.5 μL aliquot of formic acid was then added and the solution was stored in a tightly sealed container at 4°C.
Preparation of nano-particles (NPs)The NPs were prepared by adding an aqueous solution of FeCl2·4H2O (0.1 M; 20 mL) (Wako Pure Chemical Industries, Ltd.) to 20 mL of γ-aminopropyltriethoxysilane (γ-APTES) (Shin-Etsu Chemicals, Japan). After stirring for 1 h at room temperature, the solution was centrifuged at 15 krpm (Hitachi, CF15RXII) at 4°C for 1 h. The resulting precipitate was washed three times with ultrapure water.
MALDI-TOF-MSAmino acid samples (lysine and glycine) tagged with either Py0, 6 or 12 were diluted to 1, 10, 100 fmol/μL, and 1 and 10 pmol/μL. We used 2,5-dihydroxybenzoic acid (DHB) (Nacalai Tesque, Japan) (40 mg/mL) or NPs (approximately 2 mg/mL) as an ionization assisting reagent. Amino acid samples and DHB or NPs were dropped onto a target plate using a pipette. Measurements were conducted by MALDI TOF-MS (autoflex: Bruker Daltonik GmbH). The analyte surface was irradiated by 500 laser shots and MALDI TOF spectra were acquired in the positive ion detection mode.
All analyses were performed in triplicate using Nano-PALDI and MALDI MS. We detected triplet signals which correlated with Py-Tag-labelled lysine (m/z 321.2, 327.2 and 333.2) and the intervals between signals was 6 u from multiple signals, indicating successful modification by Py0, 6 and 12, respectively. The detected signal intensity for lysine by Nano-PALDI showed less noise and a higher intensity than that of MALDI (Fig. 3). Compared to the signal-to-noise ratio (S/N) from the MS signals, Nano-PALDI showed a 16 times higher intensity than the MALDI signals and a significant difference (p<0.01) was observed. In addition, Py-Tag could facilitate the measurements due to the mass shift to the positive region and can be used for comparative analyses of the same target molecules in different samples.
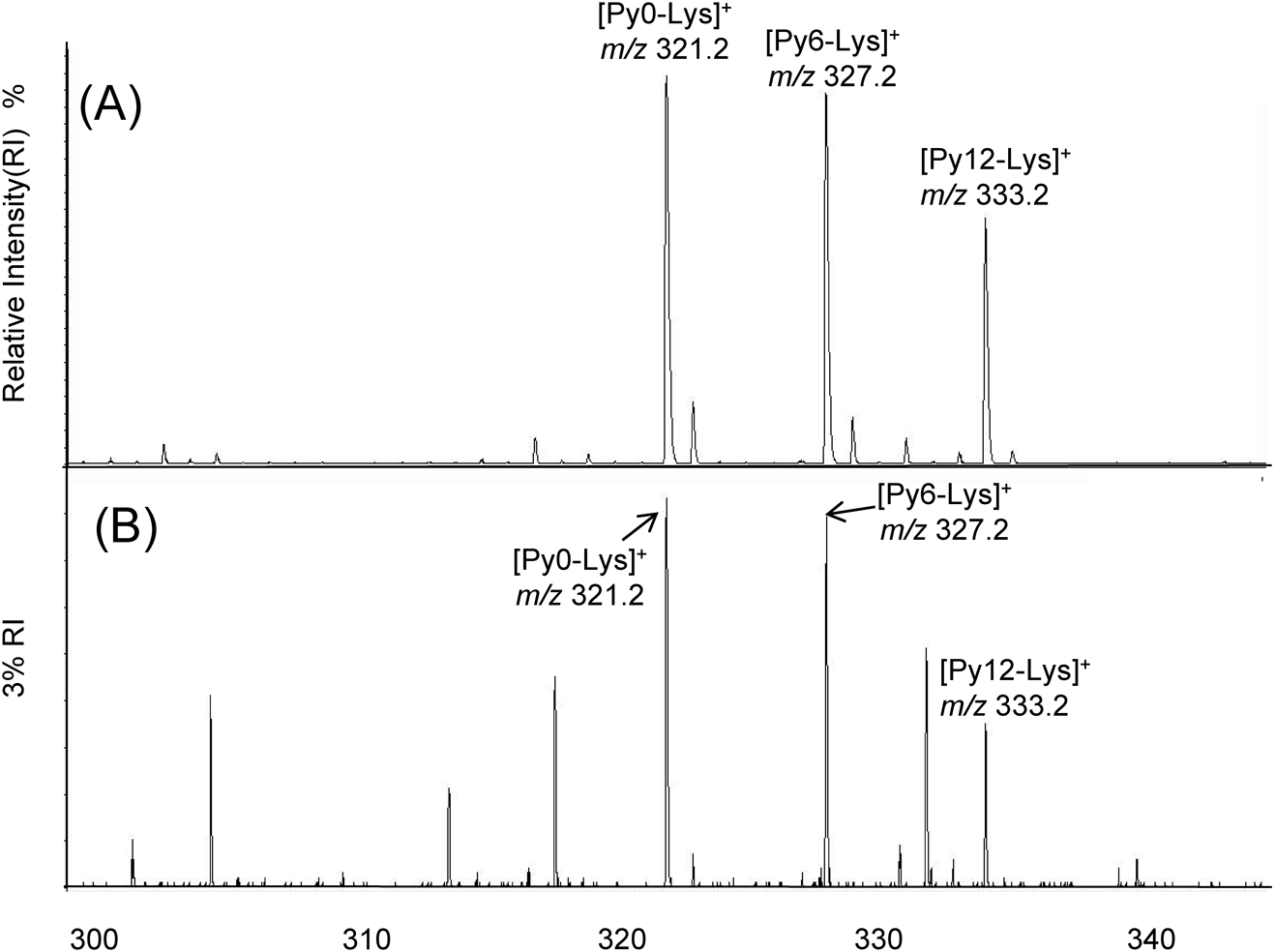
We also confirmed the LOD (limit of detection) for lysine (Table 1). The LOD for lysine calculated from the S/N obtained using Flex Analysis 3.4 software was 0.3 fmol by Nano-PALDI and 3 fmol by MALDI MS. We compared the linearity (R2) of the semilog graph for each intensity. A higher linearity was obtained by Nano-PALDI MS than by MALDI. In these studies, Nano-PALDI showed a high ionization efficiency at the femtomole level, thus making it possible to detect lower concentrations of analytes using the Nano-PALDI method.
| Method | Derivatized tag | Precursor ion (m/z) | Product ion (m/z) | Linearity (R2) | LOD (fmol) |
|---|---|---|---|---|---|
| Nano-PALDI | Py0 | 321.2 | 192.2 | 0.944 | 0.3 |
| Py6 | 327.2 | 198.6 | 0.945 | 0.3 | |
| Py12 | 333.2 | 204.6 | 0.901 | 0.3 | |
| MALDI | Py0 | 321.2 | 192.2 | 0.850 | 3.0 |
| Py6 | 327.2 | 198.6 | 0.845 | 3.0 | |
| Py12 | 333.2 | 204.6 | 0.782 | 3.0 |
For glycine, no signal was detected by either MALDI or Nano-PALDI MS because Py-Tag can only react with the primary amine of a compound that contains more than a two carbon chain. Thus, Py-Tags can specifically modify an amino acid with a long carbon chain such as lysine but not the primary amine of the N-terminus.
In addition, we conducted tandem MS to confirm the lysine fragments that were produced by Nano-PALDI. Py0-modified lysine regularly cleaved at the pyridine ring moiety to provide ions with m/z 192.2 (Fig. 4). Other fragment ions at m/z 130.1 and 275.3 were detected. From the Py6- and Py12-modified lysine, m/z 198.6 and 204.6 as an isotope labelled pyridine ring moiety were observed. The Nano-PALDI method could be applied to the analysis if the architectures of amino acids.
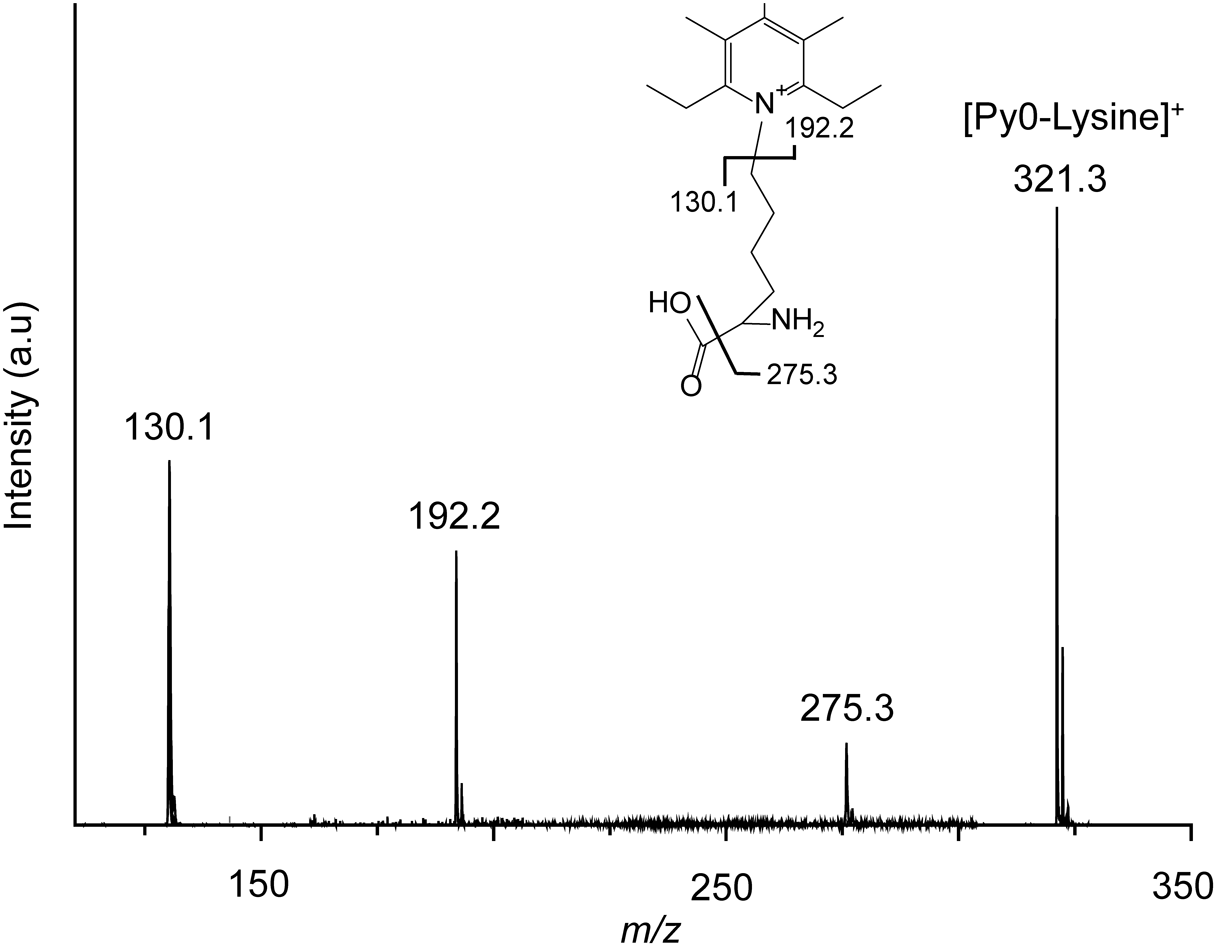
We succeeded in detecting Py-Tagged lysine by MALDI and Nano-PALDI MS. In particular, for Nano-PALDI, we obtained high intensity ions at lower concentrations of lysine and with less noise compared to MALDI MS. These results show the utility of using a combination of Nano-PALDI and Py-Tag for the analysis of amino acids.
** Py-Tag is a trademark of Taiyo Nippon Sanso Co.
We thank the members of our laboratory at FPU, particularly Ms. Y. Imase and Ms. S. Nakada. This work was supported in part by a Grant-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan (16K04890 to S.T.), The Shimadzu Science Foundation, and The Tojuro Iijima Foundation for Food Science and Technology to S. T.