-
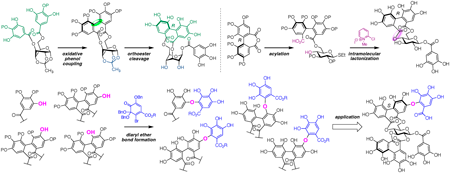
72 巻 (2024) 4 号 p. 349-359Development of Methodologies toward the Unified Synthesis of Ellagitannins もっと読む
-
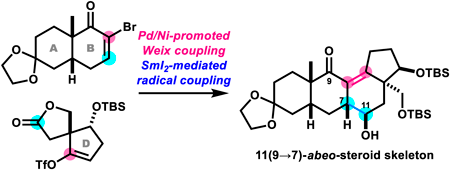
72 巻 (2024) 4 号 p. 360-364Unexpected Formation of 11(9→7)-abeo-Steroid Skeleton in Synthetic Studies toward Batrachotoxin もっと読む
-
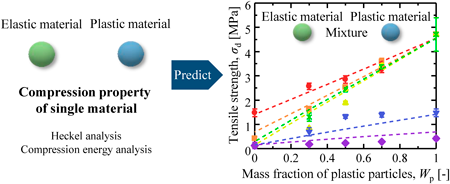
72 巻 (2024) 4 号 p. 374-380A Method for the Tensile Strength Prediction of Tablets with Differing Powder Plasticities もっと読む
-
72 巻 (2024) 4 号 p. 413-420P(III)-Mediated Formal Reductive N–H Bond Insertion Reaction of Hydrazones to α-Keto Esters もっと読む
-
72 巻 (2024) 3 号 p. 311-312Effect of Two-Photon Excitation to 8-Azacoumarin Derivatives as Photolabile Protecting Groups もっと読む
-
72 巻 (2024) 3 号 p. 336-339Synthesis of 2-Substituted Indoles via Migration Reaction of 3-Substituted Indoles with Triflic Acid もっと読む
-
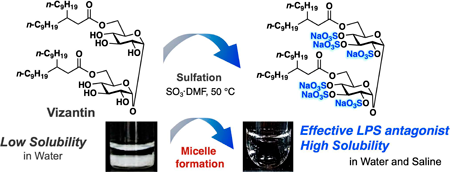
72 巻 (2024) 2 号 p. 226-233Development of a Water Soluble Self-assembling Analogue of Vizantin もっと読む
-
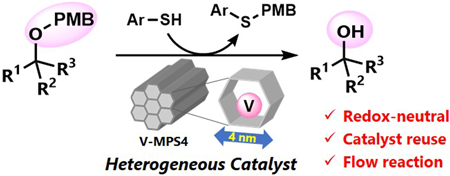
72 巻 (2024) 2 号 p. 213-219Nucleophilic Deprotection of p-Methoxybenzyl Ethers Using Heterogeneous Oxovanadium Catalyst もっと読む
-
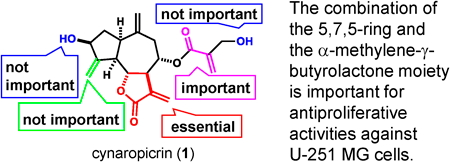
72 巻 (2024) 2 号 p. 200-208Antiproliferative Activities of Cynaropicrin and Related Compounds against Cancer Stem Cells もっと読む
-
編集者のコメント
The authors focus on the long-term consistency of dissolution profiles of generic pharmaceutical products. By analyzing a vast dataset of 1675 products across 127 ingredients, the study uncovers the intricate factors influencing changes in dissolution profiles post-approval. It emphasizes the significance of co-development in the increase of dissimilar dissolution products, the pivotal role of API particle size in poorly soluble drugs, and the impact of acidic or basic residues on dissolution changes at specific pH levels. These findings highlight the necessity for proper development that consider formulation and process variables to ensure the sustained bioequivalence of generic drugs.
-
編集者のコメント
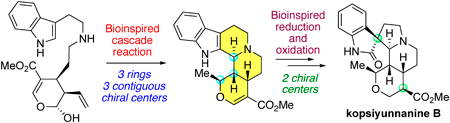
Kopsiyunnanine B was isolated from Yunnan Kopsia arborea and possesses a unique folded and complex pentacyclic structure containing six contiguous chiral centers. In this article, the authors reported the asymmetric total synthesis of Kopsiyunnanine B, along with their originally proposed biosynthetic pathway. The key transformation is an impressive cascade reaction that constructs three ring structures and three chiral centers in one step. Following the stereoselective reduction of the β-acrylate and oxidation to oxindole, the natural product is synthesized over 14 steps. Their careful consideration of the biosynthetic hypothesis has resulted in an exceptionally efficient synthesis with a minimal number of steps.
-
72 巻 (2024) 1 号 p. 80-85Five New Analogs of Streptogramin Antibiotic Viridogrisein Isolated from Streptomyces niveoruber もっと読む編集者のコメント
Given the spread of antimicrobial-resistant bacteria (AMR), there is an urgent need for the ongoing search for novel antibacterial natural products. The authors discovered five new viridogrisein congeners from Streptomyces niveoruber with potent antibacterial activities against Gram-positive bacteria. Additionally, co-treatment with griseoviridin, another natural product from the same producer, enhanced the activity. Biosynthetic studies have revealed that SgvY, encoded in the viridogrisein biosynthetic gene cluster, detoxifies viridogrisein against Staphylococcus aureus by linearization, suggesting its role in the self-resistance system in S. niveoruber. These results could facilitate the understanding of antimicrobial-resistant mechanism for developing the countermeasures against AMR.
-
72 巻 (2024) 1 号 p. 86-92Derivation of the Extended Kawakita Equation for Estimating the Yield State of Powder in Die もっと読む編集者のコメント
The Kawakita equation has been used for estimating yield pressure and porosity of compressed powder in the die. This equation assumes the compression pressure is homogeneously distributed. However, in actual powders, it is not homogeneously distributed due to the friction on the die wall. The authors extended the Kawakita equation by accounting for the inhomogeneous distribution of compression pressure. The extended Kawakita equation theoretically explained the powder behavior yielding sequentially from the loading punch to fixed punch due to the spatial limitation of particle rearrangement. Therefore, the extended Kawakita equation advances understanding of powder compaction in die.
-
編集者のコメント
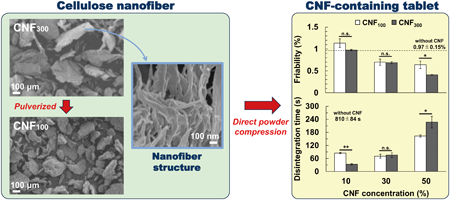
While the addition of cellulose nanofiber (CNF) to tablet formulations during direct compression has attracted increasing attention as a means for enhancing tablet strength and disintegration, they are also known to increase the variation in tablet weight and drug content. This study evaluated the effect of pulverized CNF on the variation in tablet weight and drug content. The pulverized CNF reduced both weight and drug content variation to a larger extent than untreated CNF. Further, either CNF achieved sufficient tablet strength and short disintegration time. Thus, the authors provided evidence that CNF is useful as a multifunctional additive.
-
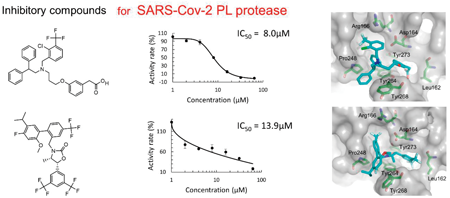
71 巻 (2023) 12 号 p. 897-905In Silico Identification of Inhibitory Compounds for SARS-Cov-2 Papain-Like Protease もっと読む編集者のコメント
Computational screening is a powerful technique for drug discovery today. In this work, a virtual screening was performed to find SARS-Cov-2 PL protease inhibitors, utilizing a chemical database consisting of approved and investigational drugs. A key issue for successful virtual screening is the accuracy of computational predictions for the binding pose and score of each compound to the target. The authors applied their original software program, Chem. Pharm. Bull., 2017, 65, 461, for calculating the score. Their approach identified five inhibitory compounds against the PL protease. The inhibitory activities were evaluated by an enzymatic assay with the FRET technique.