2019 年 42 巻 6 号 p. 977-981
2019 年 42 巻 6 号 p. 977-981
Hepatitis C virus (HCV) infection leads to chronic hepatitis, liver cirrhosis, and hepatocellular carcinoma in 50–80% of the cases. Interferons (IFNs) and the nucleoside analog ribavirin form the basis of the treatment of this infection but are not considered sufficiently effective and cause several side effects. In this study, we developed a novel viral-specific drug delivery method. Enveloped viruses, including HCV, expose an anionic phospholipid, phosphatidylserine (PS), on their surface to mediate their binding and entry into cells for infection. To target such exposed PS on HCV, we developed a chimeric recombinant protein containing human IFN and mouse lactadherin (also known as milk fat globule epidermal growth factor 8), which binds with high affinity to PS. The IFN–lactadherin fusion protein showed a high binding affinity toward PS and HCV and consequently blocked viral replication in the infected cells more efficiently than conventional IFN. Overall, these data suggest that conjugation with lactadherin facilitates the delivery of any protein drug to PS-exposing enveloped viruses.
Hepatitis C virus (HCV) is an enveloped, positive-sense, single-stranded RNA virus belonging to the hepacivirus genus of the Flaviviridae family.1) Globally, chronic infection with HCV is considered a major healthcare problem, affecting 3% of the world population, i.e., approximately 170–200 million individuals.2) Since the identification of HCV, antiviral therapy based on interferon (IFN)-α, administered alone or in conjunction with the nucleoside analog ribavirin, has formed the basis of treatment regimens.3) IFNs are a multigene family of inducible cytokines that stimulate cells via membrane receptors.4) Typically, type I IFNs are produced early upon virus infection as a first-line defense.5) These INFs induce hundreds of IFN-stimulated genes, thereby inhibiting various stages of viral replication and enhancing host response6); in addition, type I IFNs have some immunoregulatory activities.7) However, these treatments are frequently associated with various adverse effects, including influenza-like symptoms, hematological changes, and neuropsychiatric disturbances.8) Therefore, the development of a novel method in which IFN exhibits a more specific action on viruses is expected.
Several recent findings suggested that enveloped viruses expose negatively charged phospholipids such as phosphatidylserine (PS), which play a role in mediating virus entry.9–11) The presentation of PS on the outer leaflet of enveloped viruses disguises them as apoptotic bodies, thereby conning cells into engulfing the virions through cell clearance mechanisms. This mechanism of enhanced viral entry is termed as “apoptotic mimicry.” PS is generally localized to the inner leaflet of plasma membranes but is redistributed and exposed on the outer membrane when a cell undergoes apoptosis.12) The exposure of PS on apoptotic cells provides a key signal that triggers cell engulfment.13) Apoptotic mimicry was first hypothesized to be used by hepatitis B virus.14) Later, the requirement of viral PS for efficient entry and infection was proposed for viruses such as human immunodeficiency virus (HIV), Ebola, Marburg, vaccinia, and Pichinde viruses.15) Recent findings indicate that even nonenveloped viruses have evolved strategies to engage apoptotic clearance receptors for internalization.16) It has been proposed that nonenveloped viruses such as hepatitis A virus and poliovirus become enveloped in the host cell membrane when they bud into the host cell organelles, i.e., multivesicular bodies or PS vesicles.17,18)
The soluble protein Gas6 binds to PS on the virion surface and bridges viruses to the cell surface by interacting with the tyrosine kinase receptor Axl; the formation of this complex is necessary for enhancement of virus entry.19) Various additional PS receptors have been identified to date, including T-cell immunoglobulin and mucin domain TIM-1 and TIM-4 proteins and lactadherin/integrin αvβ3 or αvβ5 complexes.13,20) Reportedly, the mutation of the residues involved in PS binding or complex formation of these receptors results in the inhibition of viral infection.21) Moreover, viral internalization into HEK293T cells was significantly enhanced by the overexpression of these PS receptors and was inhibited by competing with PS liposomes,22) providing evidence that virion-associated PS receptor interactions are responsible for virion uptake, and not just for virus binding. In particular, recent studies demonstrated that TIM-1 promotes HCV infection by serving as an attachment receptor for binding to PS exposed on the HCV envelope.23) The knockout of the TIM-1 gene or use of PS-containing liposomes resulted in a remarkable reduction of HCV cell attachment and subsequent infection. The property that HCV strongly exposes PS on its surface prompted the possibility that antiviral drugs such as IFN can be specifically delivered to the virus by targeting PS. Therefore, we employed a unique soluble protein (lactadherin) that strongly binds to PS.20) In this study, we developed a chimeric protein in which human IFN was conjugated to mouse lactadherin that has a strong affinity toward PS and showed that the engineered protein can be a novel drug that efficiently prevents the replication of HCV by specifically targeting the IFN to the virus.
A chimeric gene was constructed using recombinant PCR to express a fusion protein comprising N-terminal human IFN-β and C-terminal mouse lactadherin. Synthesized human IFN-β cDNA (FASMAC, Japan) was amplified by PCR using the following primer pair: IFN-forward (5′-ACC ATG ACC AAC AAG TGT CTC CT-3′) and IFN-reverse (5′-GGA GTC ACA GAA GTC GTT TCG GAG GTA ACC-3′; the lactadherin sequence is underlined). Mouse lactadherin cDNA coupled to the FLAG-tag coding sequence was obtained by PCR using the pCAG–lactadherin (D89E) plasmid as a template in which the FLAG-tag sequence was conjugated to the 3′-end and an amino acid substitution was introduced in the RGD motif to inactivate phagocytosis, as described previously,20) with the following primer pair: lactadherin-forward (5′-GGT TAC CTC CGA AAC GAC TTC TGT GAC TCC-3′; the IFN sequence is underlined) and lactadherin-reverse (5′-TTA CTT GTC GTC GTC GTC CTT GT-3′). Both PCR products were purified, mixed, and further amplified by the second PCR with the IFN-forward and lactadherin-reverse primers. Then, the chimeric IFN–lactadherin cDNA was inserted into the pCAG vector.
Purification of Lactadherin or the IFN–Lactadherin Fusion ProteinpCAG–lactadherin (D89E) or pCAG–IFN–lactadherin was transfected in HEK293T cells using polyethylenimine (PEI); PEI/DNA complexes were formed using a 10 : 1 PEI amine:DNA mixture by diluting 20 µg DNA in a 2 mL reduced serum medium (Opti-MEM, Gibco, U.S.A.). The cell-conditioned medium was collected, and the supernatant was cleared from the cells and debris by filtration through a 0.2-µm flow filter unit (Thermo Fisher, U.S.A.). The FLAG-tagged lactadherin or IFN–lactadherin fusion protein was captured through column chromatography (Bio-Rad, U.S.A.) using anti-FLAG M2 affinity gel (Sigma, U.S.A.) according to the manufacturer’s protocol. Briefly, the supernatants were passed through a 200-µL resin column by gravity flow, and the resin was washed with 20 column volumes of phosphate-buffered saline (PBS) to remove any proteins unbound to the M2 antibody. The FLAG peptide (100 µg/mL; Sigma) was used for the competitive elution of the C-terminal FLAG-tagged fusion protein. Next, the eluate was concentrated with Amicon Ultra centrifugal filter 10K (Millipore, Germany) to remove the residual FLAG peptide. The quantity and purity of the protein were estimated by sodium docedyl sulfate-polyacrylamide gel electrophoresis followed by visualizing with Oriole Fluorescent Gel Stain (Pierce, U.S.A.).
Enzyme-Linked Immunosorbent Assay (ELISA) for PS BindingSolid-phase ELISA for PS binding was conducted as described previously.24) In brief, PS was diluted to 500 µM in methanol; 100 µL of the solution was used to coat a 96-well microplate (Fisherbrand) and air-dried at room temperature. Protein-binding sites in the coated well were blocked with 10 mg/mL bovine serum albumin for 3 h. Lactadherin or IFN–lactadherin fusion protein was serially diluted to the concentrations 30, 20, 10, 5, 2.5, 1.25, and 0.625 nM in PBS containing 0.05% Tween 20 (PBST); 50 µL of the protein solution was added to the wells. The wells were incubated for 1 h at room temperature and then washed thrice with PBST. Lactadherin or IFN–lactadherin bound to the wells was quantified by ELISA with biotinylated anti-FLAG M2 antibody (Sigma), horseradish peroxidase-conjugated streptavidin, and a peroxidase detection kit (ELISA POD TMB, Nacalai Tesque, Japan). The absorbance of each sample was quantified at dual wavelengths of 450 and 650 nm using a microplate reader (iMark, Bio-Rad).
HCV Binding AssayThe binding ability to HCV was compared between the IFN–lactadherin fusion protein and recombinant human IFN-β protein (rhIFN-β; Wako, Japan) as follows: 2 µg proteins was used as the input and incubated with 1 × 105 IU HCV virions for 1 h at 4°C. The virus–protein mixture was passed through a 100 K centrifugal filter (Amicon, Millipore) and washed twice with PBS to eliminate unbound proteins. HCV virions were recovered and subjected to Western blot analysis using anti-human IFN-β antibody (BioLegend). Then, the signal ratio of HCV-bound proteins per input proteins was compared using the ImageQuant LAS 4000 Luminescent Image Analyzer (GE Healthcare, U.S.A.).
HCV Infection AssayHCV virions (1.5 × 104 IU) were incubated with 400 nM (or 100 and 25 nM) IFN–lactadherin or rhIFN-β in 100 µL solutions for 1 h at 4°C. To remove unbound proteins, the virus–protein mixture was passed through a 100 K centrifugal filter and washed twice with PBS. The mixture was added to the human hepatoma cell line Huh-7.5.1 plated at a density of 3 × 104 cells per well in a 24-well plate in Dulbecco’s modified Eagle’s medium (DMEM) containing 10% fetal bovine serum at 37°C (multiplicity of infection = 0.5). The infection was allowed to proceed for 2 h; then, the cells were washed with PBS and supplemented with fresh complete DMEM. At postinfection 2 d, total RNA was isolated from the infected cells, and the levels of HCV infection were determined by RT-quantitative PCR (QPCR). Total cellular RNA was isolated using the GenElute Mammalian Total RNA Miniprep Kit (Sigma) according to the manufacturer’s instructions. RT reaction was performed using the High-Capacity RNA-to-cDNA Kit (Applied Biosystems). Further, QPCR analysis was performed using the SYBR qPCR Mix from Toyobo, Japan and the Roche LightCycler 96 system. HCV and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) transcript levels were quantified using the following primer pairs, respectively: HCV-forward (5′-GAG TGT CGT GCA GCC TCC A-3′) and HCV-reverse (5′-CAC TCG CAA GCA CCC TAT CA-3′) and GAPDH-forward (5′-TGT AGT TGA GGT CAA TGA AGG G-3′) and GAPDH-reverse (5′-ACA TCG CTC AGA CAC CAT G-3′). Data were analyzed using the ΔCt method and normalized against the levels of GAPDH RNA expression in each sample.
Lactadherin, or milk fat globule epidermal growth factor (EGF) 8 (MFG-E8), is a glycosylated protein that was first discovered as a component of milk fat globule membranes.25) Mouse lactadherin contains two EGF domains: proline/threonine-rich domain (P/T-rich domain) and two factor VIII-homologous domains (C-1, C-2; Fig. 1). Lactadherin binds to PS exposed on apoptotic cells, extracellular vesicles, and enveloped viruses via C-1 and C-2.26) Using this property, we developed a chimeric fusion protein in which mouse lactadherin was connected to human IFN-β. We used mouse lactadherin because the P/T-rich domain, which is absent in human lactadherin, contributes to affinity enhancement toward PS by 10 times.20) Because the RGD motif in the second EGF domain promotes phagocytosis by binding to the integrin αvβ3 or αvβ5 on phagocytes, we introduced an amino acid substitution from D to E in this motif. The protein is tagged with FLAG sequence in the C-terminus for affinity purification.
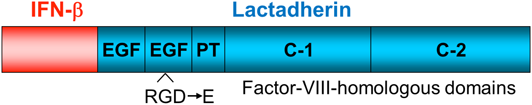
Mouse lactadherin (blue), a 75-kDa secreted protein that binds to PS via its C-1 and C-2 domains, was fused to human IFN-β (red). An amino acid substitution (from D to E) was introduced to the RGD motif in the second EGF domain that mediates the interaction with the integrin receptors present on phagocytes. (Color figure can be accessed in the online version.)
First, to examine the binding ability of IFN–lactadherin to PS, microtiter plates were coated with PS and incubated with lactadherin or IFN–lactadherin. As shown in Fig. 2, IFN–lactadherin bound to PS in a dose-dependent manner, which is comparable to lactadherin. We found that both proteins bound to PS with a dissociation constant (Kd) of 3 nM, indicating that the recombinant IFN–lactadherin fusion protein has an affinity toward PS as strong as lactadherin.
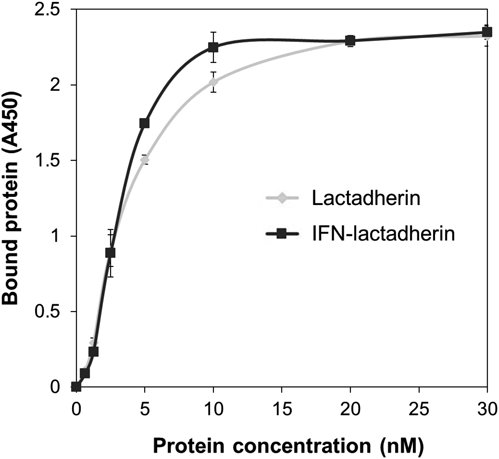
Microtiter plates coated with PS were incubated with lactadherin (diamonds) or IFN–lactadherin (squares), and the protein bound to the wells was quantified by ELISA. Assays were performed in triplicates, and the average and standard deviation values are plotted. The proteins did not bind to the plates without PS-coating.
To examine whether IFN–lactadherin binds to enveloped viruses, binding efficiency to HCV was compared between IFN–lactadherin and rhIFN-β. The virus was incubated with IFN–lactadherin or rhIFN-β and then washed with PBS. Virus-bound proteins were detected using anti-IFN-β antibody by Western blotting (Fig. 3). We found that only IFN–lactadherin bound to the virus due to the ability of lactadherin to bind to PS exposed on the viral envelope.
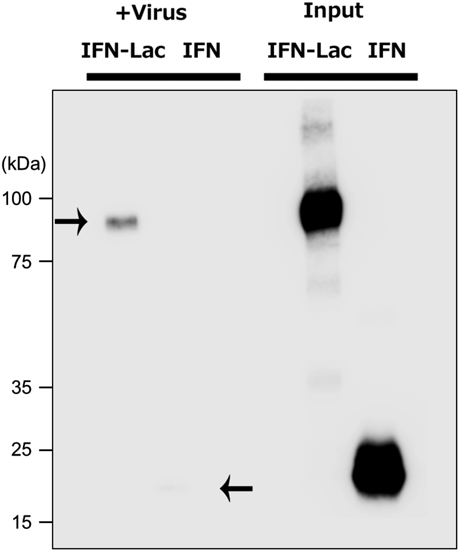
IFN–lactadherin (IFN–Lac) or IFN-β (IFN) was incubated with HCV virions (+Virus), and unbound proteins were eliminated by ultrafiltration. The proteins bound to HCV were detected by Western blotting with antibody against IFN-β (arrows). The amount of proteins used as input is shown in the right lanes.
Based on the observation that IFN–lactadherin efficiently bound to HCV, we tested the effect of inhibiting virus replication. Serially diluted IFN–lactadherin or rhIFN-β protein was incubated with HCV. The virus–protein mixture was then added to hepatoma Huh-7.5.1 cells to check the effect of both proteins on viral replication. The results showed that incubation with IFN–lactadherin or rhIFN-β at a concentration of 400 nM substantially inhibited the replication of HCV at the same efficacy as quantified by QPCR (Fig. 4). However, when unbound proteins were washed away from the virus–protein mixture, the inhibition of viral replication by rhIFN-β was markedly impaired, whereas that by IFN–lactadherin remained unchanged. Moreover, even a lower concentration of IFN–lactadherin (100 and 25 nM) with a washing away step still retained the inhibitory ability, comparable to higher concentration (400 nM) due to the binding to HCV.
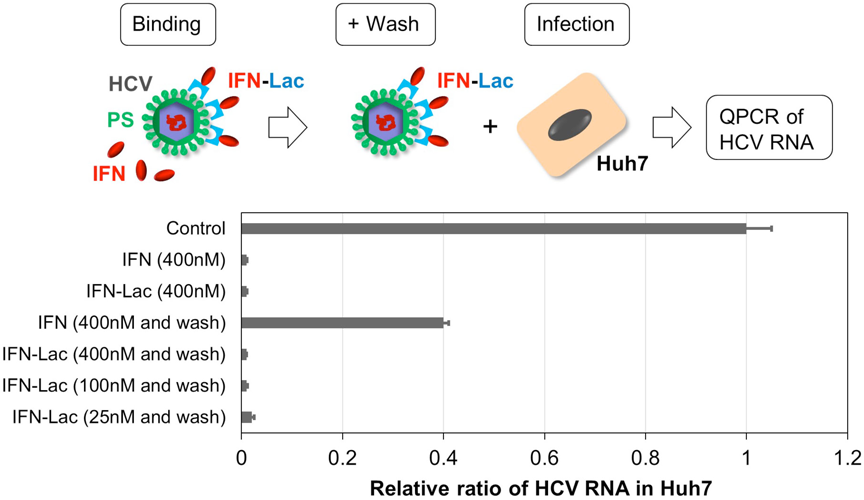
HCV virions were incubated in the absence (Control) or presence of the indicated concentrations of IFN or IFN–Lac for 1 h. The virus–protein mixture with or without the elimination of unbound proteins by washing with PBS was added to Huh-7.5.1 cells, and the levels of HCV RNA normalized to GAPDH in the cells were determined by QPCR at postinfection 2 d. The relative ratio to control is shown. Experiments were performed in triplicates, and the average and standard deviation values are plotted. (Color figure can be accessed in the online version.)
In summary, our data indicated that IFN–lactadherin bound to HCV with a high affinity, which is not characteristic to conventional IFN. We initially thought that lactadherin would inhibit the uptake of viruses by masking PS on viral membrane, but even a high concentration of lactadherin did not inhibit the HCV infection (data not shown). Therefore, we employed the PS binding property of lactadherin to develop this drug delivery method to enveloped viruses. In this study, we washed viruses to remove unbound proteins from the virus–protein mixture. This process is intended to mimic the blood flow in the human body, assuming that blood flow washes IFN away from healthy organs (which are absent in viruses) and delivers IFN only to the site harboring viruses. The used lactadherin-conjugation method could be applied for specific delivery of any protein drug to enveloped viruses by targeting the PS exposed on their surface, which may provide a novel strategy for inhibiting infection by many viruses that use apoptotic mimicry and contribute to reduce the side effects caused by the drug.
We thank Drs. Yoshiharu Matsuura and Toru Okamoto (Osaka University) for providing the HCV virions. This work was supported by the AMED Research Program on Hepatitis to R.H. (H24-013) and by the World Premier International Research Center Initiative (WPI), Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan.
The authors declare no conflict of interest.