- 1 号 p. 1-
- |<
- <
- 1
- >
- >|
-
渡邉 力也2024 年 64 巻 1 号 p. 1
発行日: 2024年
公開日: 2024/03/25
ジャーナル フリー HTMLPDF形式でダウンロード (129K) HTML形式で全画面表示
-
ジャーナル フリー HTML
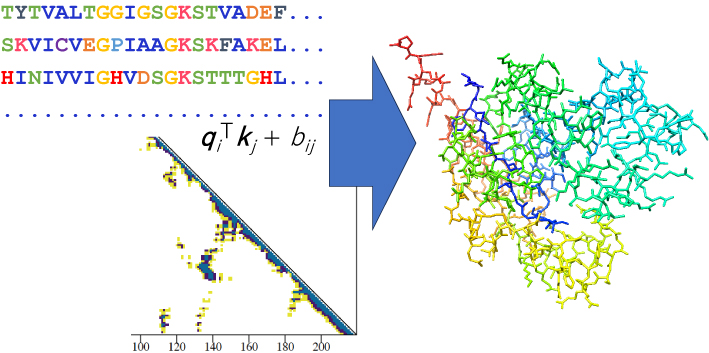
電子付録大量の蛋白質配列・構造データに基づく深層学習モデルであるAlphaFoldによって,多くの蛋白質の高精度な構造予測モデルが利用可能となっている.本稿では,AlphaFold/AlphaFold-Multimerを用いた蛋白質構造/複合体構造予測の背景や効果,利用の際の注意点などについて概説する.
 抄録全体を表示PDF形式でダウンロード (884K) HTML形式で全画面表示
抄録全体を表示PDF形式でダウンロード (884K) HTML形式で全画面表示
-
ジャーナル フリー HTML
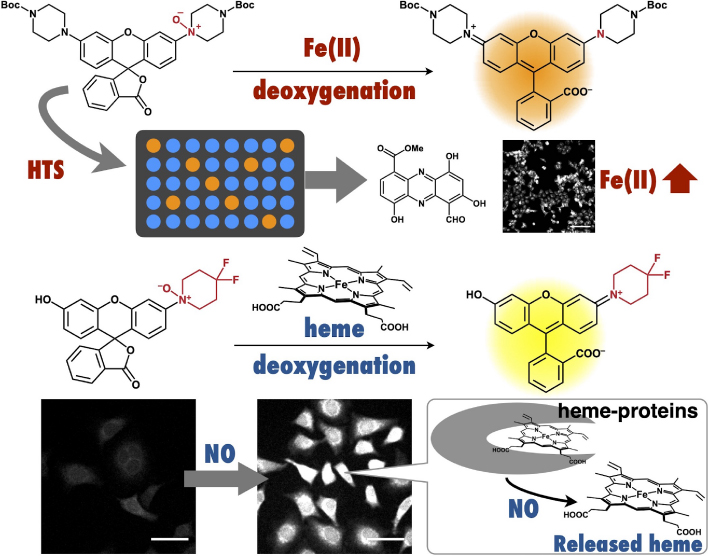
鉄は地球上に確認されているあらゆる生物に含まれ,鉄イオンあるいはヘムの形で多くの生命維持に必須の機能を担っていることが知られているが,細胞内における鉄イオン・ヘム自体の動態は未解明である.本総説では,これまでに筆者が開発してきた,遊離鉄イオン・遊離ヘムの生細胞イメージングプローブについて紹介する.
 抄録全体を表示PDF形式でダウンロード (1174K) HTML形式で全画面表示
抄録全体を表示PDF形式でダウンロード (1174K) HTML形式で全画面表示
-
ジャーナル フリー HTML
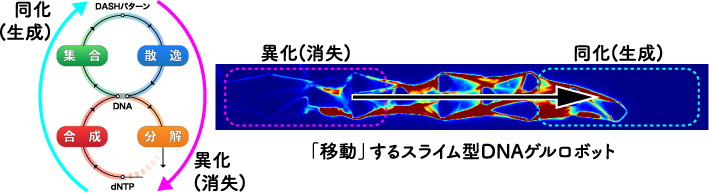
近年,DNAを使った新たな構造作製手法として「DNAハイドロゲル」が注目されている.我々は,酵素反応で作るDNA物理ゲルの技術をもとにして,代謝を模したDNA材料システム「DASH」を開発,また,これを使って駆動する「スライム型分子ロボット」の原型を作り出した.本トピックスでは,このシステムの概要と展望を述べる.
 抄録全体を表示PDF形式でダウンロード (2139K) HTML形式で全画面表示
抄録全体を表示PDF形式でダウンロード (2139K) HTML形式で全画面表示 -
森田 陸離, 重田 育照, 原田 隆平2024 年 64 巻 1 号 p. 21-24
発行日: 2024年
公開日: 2024/03/25
ジャーナル フリー HTMLドッキングシミュレーションは化合物とタンパク質の複合体構造を高速に予測する計算手法である.しかし,膜タンパク質について脂質膜の存在をあらわに考慮していないという問題があった.我々は分子の疎水性指標であるlogPを用いてドッキングスコアを補正する手法(LoCoMock)を考案した.
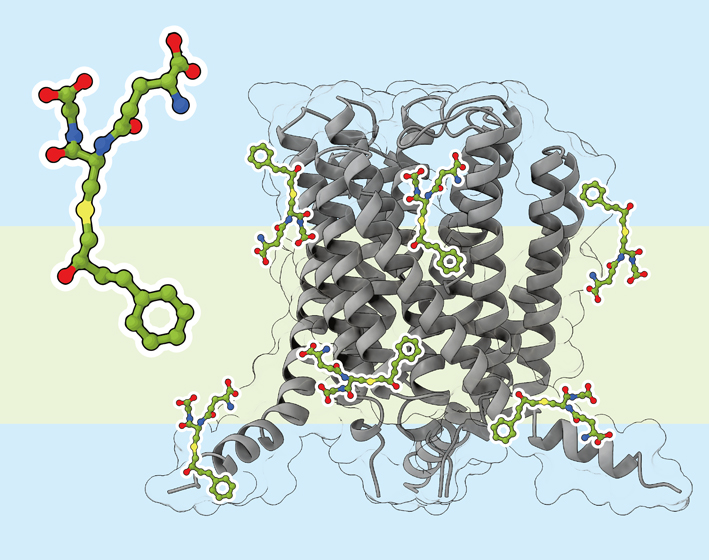 抄録全体を表示PDF形式でダウンロード (1965K) HTML形式で全画面表示
抄録全体を表示PDF形式でダウンロード (1965K) HTML形式で全画面表示 -
山下 敦子, 岡崎 圭一2024 年 64 巻 1 号 p. 25-27
発行日: 2024年
公開日: 2024/03/25
ジャーナル フリー HTML腸内シュウ酸分解菌は,私たちが摂取したシュウ酸を吸収し,代謝分解してギ酸として排出することで,宿主の尿路結石症発症リスクを低減する.この過程の鍵を握るのが,分解菌細胞膜に存在するシュウ酸/ギ酸対向輸送体OxlTである.本稿では,筆者らが解析したOxlTの構造と構造動態から,その分子メカニズムを概説する.
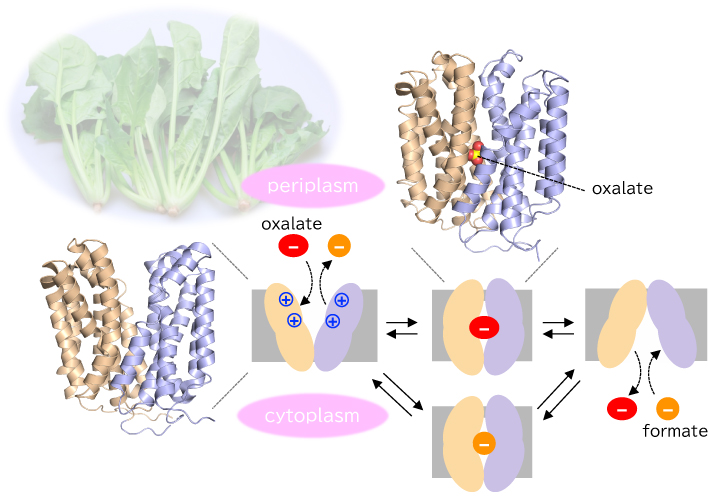 抄録全体を表示PDF形式でダウンロード (3053K) HTML形式で全画面表示
抄録全体を表示PDF形式でダウンロード (3053K) HTML形式で全画面表示 -
赤井 大夢, 小岩 滉宜, 井澤 幸広, 庄司 観2024 年 64 巻 1 号 p. 28-31
発行日: 2024年
公開日: 2024/03/25
ジャーナル フリー HTMLDNAナノチャネルは,緻密な配列設計や化学修飾により,チャネルタンパク質の機能模倣に成功している一方で,その膜挿入効率は低いため,応用展開は制限されている.本稿では,DNAナノチャネルを物理的に膜挿入することで,膜挿入効率を従来の3倍に向上させた「DNAナノチャネルプローブ技術」について紹介する.
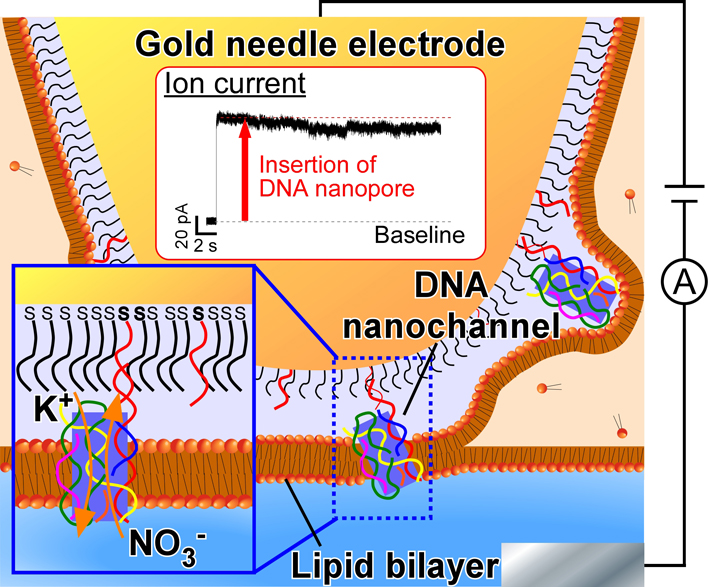 抄録全体を表示PDF形式でダウンロード (2123K) HTML形式で全画面表示
抄録全体を表示PDF形式でダウンロード (2123K) HTML形式で全画面表示 -
ジャーナル フリー HTML
動物オプシンは,三量体Gタンパク質を介するシグナル伝達経路を光で操作するツールとして活用されている.しかし,三量体Gタンパク質が複数のシグナル経路を駆動するために,何を光操作しているのか不明確になる難点があった.この難点を(部分的に)克服できる結果を得られたので,本トピックスで紹介する.
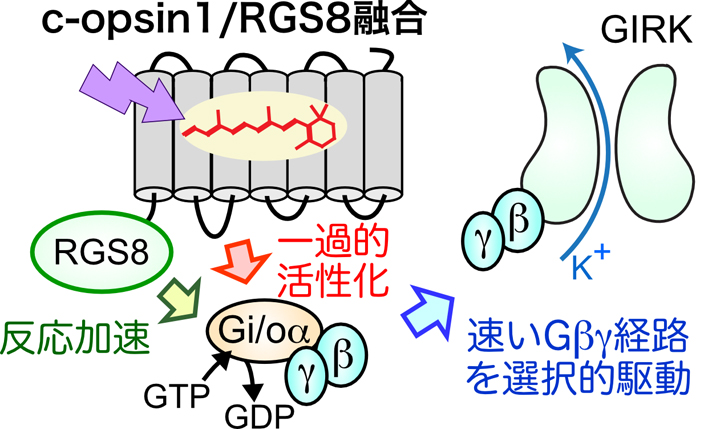 抄録全体を表示PDF形式でダウンロード (962K) HTML形式で全画面表示
抄録全体を表示PDF形式でダウンロード (962K) HTML形式で全画面表示 -
ジャーナル フリー HTMLPDF形式でダウンロード (1547K) HTML形式で全画面表示
-
西出 亮介, 石原 秀至2024 年 64 巻 1 号 p. 38-41
発行日: 2024年
公開日: 2024/03/25
ジャーナル フリー HTML生物に見られるパターンの多くは曲面上で生じる.曲面はパターンダイナミクスを変化させ,生物学的な役割を担うことがわかってきた.私達は,曲面上でTuringパターンの振る舞いを調べることで,曲率がパターンを動かす効果を持っていることを明らかにした.平面上でパターンが止まっていても,曲面上でもそうとは限らない.
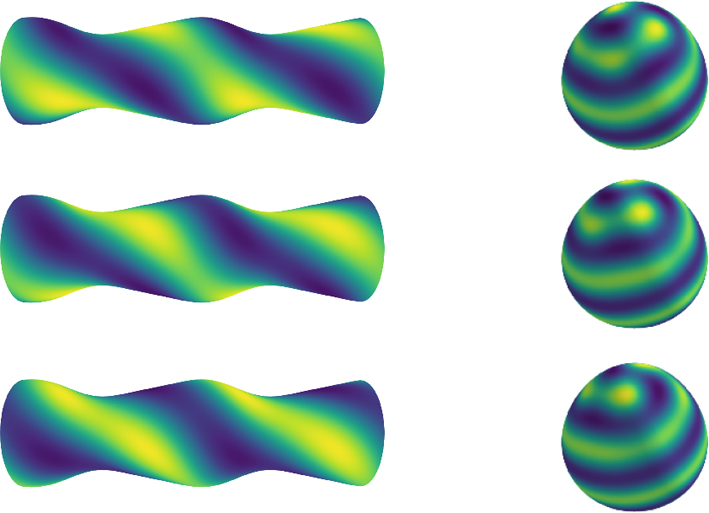 抄録全体を表示PDF形式でダウンロード (1829K) HTML形式で全画面表示
抄録全体を表示PDF形式でダウンロード (1829K) HTML形式で全画面表示
-
Chun-Biu LI2024 年 64 巻 1 号 p. 44-46
発行日: 2024年
公開日: 2024/03/25
ジャーナル フリー HTMLPDF形式でダウンロード (523K) HTML形式で全画面表示
-
~第14回 中国四国支部大会~冨永 貴志2024 年 64 巻 1 号 p. 47-48
発行日: 2024年
公開日: 2024/03/25
ジャーナル フリー HTMLPDF形式でダウンロード (1932K) HTML形式で全画面表示
-
~第63回生物物理若手の会夏の学校開催報告~高橋 大地2024 年 64 巻 1 号 p. 49-50
発行日: 2024年
公開日: 2024/03/25
ジャーナル フリー HTMLPDF形式でダウンロード (1085K) HTML形式で全画面表示
-
~助けられっぱなしの海外滞在記~本田 信吾2024 年 64 巻 1 号 p. 51-52
発行日: 2024年
公開日: 2024/03/25
ジャーナル フリー HTMLPDF形式でダウンロード (1573K) HTML形式で全画面表示
-
2024 年 64 巻 1 号 p. 42-43
発行日: 2024年
公開日: 2024/03/25
ジャーナル フリー HTMLPDF形式でダウンロード (162K) HTML形式で全画面表示
- |<
- <
- 1
- >
- >|