
- |<
- <
- 1
- >
- >|
-
Yuki Kondo, Atsuki Suguchi, Hiroo Fukuda2017 Volume 82 Issue 4 Pages 335
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSDownload PDF (904K)
-
Sachihiro Matsunaga, Tomoko M. Matsunaga2017 Volume 82 Issue 4 Pages 337-339
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSA padlock probe is a circularized oligonucleotide containing complementary sequences of target regions. Padlock probes combined with rolling-circle amplification enable fluorescence in situ hybridization (FISH) to reproducibly detect the genomic position of low or single-copy DNA sequences. The FISH with padlock probes revealed that a plant gene cluster of tandemly arrayed three paralogues responsible for photosynthesis rapidly moved from the interior subnuclear region to the peripheral subnuclear region in response to light. Using a pair of peptide nucleic acids as a double-strand opener for the target region, the efficiency of FISH with padlock probes at the detection of a single-copy genomic region was much improved. FISH with padlock probes will give us reliable data of dynamics, arrangement and position of a specific single locus in both the nucleus and the condensed chromosomes.
View full abstractDownload PDF (402K)
-
Noriko Inada2017 Volume 82 Issue 4 Pages 341-348
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSObligate biotrophic microorganisms, both symbiotic and pathogenic, colonize only living host plants to complete their life cycles. Colonization of obligate biotrophs induces significant reorganization of plant cell architecture, including cytoskeletons. The reorganization of cytoskeletons, which includes microtubules (MTs) and actin microfilaments (AFs), has been extensively documented for interactions with host plants and symbiotic biotrophs, nodule-forming rhizobia, and arbuscular mycorrhizal fungi. Recent studies have identified host factors that are involved in the reorganization of AFs and that function in support of colonization of nodule-forming rhizobia. For the cytoskeletal reorganization induced by pathogenic biotrophs, we recently reported the dynamic changes of host AFs during colonization of fungus causing powdery mildew disease. This review summarizes recent advances in studies of plant cytoskeletons and their role in interactions between plants and obligate biotrophs.
View full abstractDownload PDF (596K)
-
Sukjai Rattanayuvakorn, Alongklod Tanomtong, Sumalee Phimphan, Wiwat S ...2017 Volume 82 Issue 4 Pages 349-354
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSThe present study was undertaken to identify whether any difference exists in karyotype between the tusker (chang Phai) and tuskless (chang Sridor) male Asian elephants (Elephas maximus, Linn. 1758). The elephant using in the present work are owned of some local people who lived in the elephant village, Surin Province, Thailand. Peripheral bloods were collected from the ear-vein of healthy elephants. The lymphocyte culture and harvesting method was employed follow standard protocols. Conventional, GTG-, and Ag-NORs banding techniques were applied to find out the different of both elephant karyotypes. The results showed that the standardized karyotype of both elephants were consisted of one pair of large acrocentric, four pairs of large telocentric, one pair of medium submetacentric, one pair of medium acrocentric, six pairs of medium telocentric, two pairs of small metacentric, two pairs of small submetacentric, and ten pairs of small telocentric homologous chromosomes. The X sex-chromosome was a medium metacentric and the Y sex-chromosome was a small acrocentric chromosome. Here, the NORs bearing chromosome were found on one pair of small submetacentric and two pairs of small telocentric chromosomes similar in both of male elephants. However, those classical cytogenetic techniques could not be distinguished the karyotype of tusker and tuskless male elephants. At least, we present the standardized karyotype, idiogram and the basic cytogenetic data that could be served as a basis for future chromosome analyses of Asian elephant populations and for comparative cytogenetic studies with other related species. The karyotype formula could be deduced as:
 View full abstractDownload PDF (1253K)
View full abstractDownload PDF (1253K) -
Sanjeev Kumar, Santosh Kumari, Raghbir Chand Gupta, Vikas Kumar Sharma2017 Volume 82 Issue 4 Pages 355-361
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSAt present, population-based meiotic studies were carried out on six species of genus Clematis for documenting the genetic diversity from selected localities of Himachal Pradesh in the Western Himalayas (India). All the populations are found to be diploid with meiotic chromosome number as 2n=16. Impaired male meiosis was noted in all populations of C. buchananiana, C. connata, C. gouriana, C. grata, C. montana, and C. orientalis. However, male meiosis shows variation in meiotic behaviour in the different populations. The presence of B chromosomes in C. orientalis is reported for the first time, previously, the species is also known to have 2n=16 but without B chromosomes. These anomalous species were marked with meiotic abnormalities in the form of cytomixis, chromosomal stickiness, unoriented bivalents, formation of laggards and bridges resulting in abnormal microsporogenesis, and production of heterogeneous-sized fertile pollen grains along with reduced pollen fertility.
View full abstractDownload PDF (1480K) -
Keylla Souza dos Santos, Adriana Rodrigues Passos, Janay Almeida dos S ...2017 Volume 82 Issue 4 Pages 363-367
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSThe present study aims to describe the behavior of the chromosomes in the microsporogenesis and to estimate the pollen viability in Physalis ixocarpa, for purposes of genetic improvement of the species. Floral buds previously collected and fixed in Carnoy (3 : 1) were analyzed under a microscope after being stained with acetic carmine (1%). The number of cells present in each phase of meiosis I and II were determined: prophase, metaphase, anaphase, telophase and the formation of tetrads. The irregularities observed in each of the analyzed phases were recorded. The Meiotic Index (MI) was calculated. The viability of the pollen was determined according to the level of staining: pollen stained red as viable and yellowish-green or colorless as non-viable. The percentage of viable pollen grains was determined by the number of pollen grains stained by the total number of evaluated pollen ×100. A total of 2256 meiocytes were observed. Of these, 788 meiocytes were measured in meiosis I, 762 cells were observed in meiosis II and 706 tetrads. The calculated MI for P. ixocarpa was 100%, indicating that the species are meiotically stable. The estimated pollen viability for the evaluation of 3600 pollen grains was 93.11% and considered highly viable. In the present study, the presence of pollenkitt adhered to the pollen grains was observed. Thus, the microsporogenesis process is normal, resulting in the formation of highly viable pollen grains.
View full abstractDownload PDF (543K) -
Syed Mudassir Jeelani, M. A. A. Siddique, Umer Farooq2017 Volume 82 Issue 4 Pages 369-374
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSThe present study was performed for the Himalayan medicinal species Sium latijugam on male meiosis, morphological parameters and distribution pattern of diploid as well as tetraploid cytotypes. Both diploid (2n=12) and tetraploid (2n=24) cytotypes were found in Kashmir, however, the tetraploid cytotype is new addition to the species for the first time. The tetraploids were restricted to higher altitudes and characterized by abnormal meiosis leading to high pollen sterility and formation of variable sized pollen grains. Additionally tetraploids were morphologically more luxuriant in comparison to their diploid counterparts.
View full abstractDownload PDF (578K) -
Maninder Kaur, Himshikha, Vijay Kumar Singhal2017 Volume 82 Issue 4 Pages 375-384
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSPresent study records the existence of tetraploid and hexaploid meiocytes and subsequently the formation of large and very large-sized pollen grains for the first time in the diploid individuals of Achillea millefolium. In majority of the diploid accessions, meiocytes exhibited nine bivalents, equal segregation of chromosomes during anaphases, regular tetrads and normal-sized ‘n’ pollen grain formation. In the accessions scored from Palchan (2400 m) and Dhundhi (3050 m) regions of Solang Valley, two to three proximate PMCs fused during early stages of Prophase-1 and Metaphase-1 resulting into tetraploid (4x) and hexaploid (6x) meiocytes with 18 and 27 bivalents, respectively. Although the frequency of polyploid PMCs was rather low these are detectable due to their larger size compared to diploid PMCs. In both the accessions, the diploid and polyploid PMCs exhibited the multiple chromosomal associations of four to eight chromosomes indicating the existence of structural heterozygosity. Analysis reveals that there is an increase in the chiasma frequency with structural heterozygosity for reciprocal translocations. Syncyte meiocytes followed a regular meiotic course resulting into the formation of normal tetrads but the products of such sporads yielded atypical (large and very large-sized) pollen grains compared to normal sized typical pollen grains. Although the exact cytological status of aforesaid pollen grains could not be ascertained but such fertile larger-sized pollen grains with possible ‘2n’ or ‘3n’ genetic constitution might be involved in fertilization to generate polyploid offsprings and the origin of intraspecific polyploid cytotypes (4x, 6x) which are known to exist in some populations of Himalayas in India.
View full abstractDownload PDF (1256K) -
Micheli Sossai Spadeto, Tatiana Tavares Carrijo, Carlos Roberto Carval ...2017 Volume 82 Issue 4 Pages 385-390
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSMyrsine coriacea is a Primulaceae species considered ecologically important for colonizing degraded areas and providing fruits for birds. This species has been gaining attention at present due to possessing pharmacological compounds explored in cancer treatment. This study aimed to establish the first procedure for in vitro propagation of M. coriacea through seed germination and somatic embryogenesis. A high frequency of germination (100%) was observed from mature seeds pre-treated with hydrochloric acid and inoculated into medium supplemented with gibberellic acid. Plantlets were generated after 41 days from these seeds, a time relatively short in comparison to ex vitro methods used for propagation of this species. Besides the direct system starting with seed germination, the recovery of M. coriacea was established from indirect somatic embryogenesis using immature zygotic embryos. From these explants, friable calli were formed on medium supplemented with 6-benzylaminopurine, and somatic embryos were regenerated and plantlets recovered after 115 days. This result evidenced that the immature zygotic embryos of M. coriacea possess levels of endogenous phytohormones, mainly auxins, which are sufficient to trigger the embryogenic system. All M. coriacea plantlets showed the same nuclear 2C value and chromosome number in relation to explant donors. Thus, no karyotype change occurred during propagation in the in vitro system standardized. The first reliable and relatively rapid procedures provided morphologically normal seedlings of M. coriacea, providing the basis for other tissue culture applications. Furthermore, the plantlets regenerated can be used in reforestation and conservation programs, and for production of pharmacological compounds.
View full abstractDownload PDF (491K) -
Kamlesh Kumari, Manjit Inder Singh Saggoo2017 Volume 82 Issue 4 Pages 391-394
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSPolygonatum cirrhifolium commonly known as Sobnyam in Kinnaur is rich source of starch, pectin, protein and fiber content. In the present study, two accessions of P. cirrhifolium belonging to family Asparagaceae were studied for male meiosis from the cold desert region of Kinnaur. Both the accessions showed the presence of 16 bivalents (based on x=8) at diakinesis and metaphase, which is in conformity with the previous reports. Earlier, floristic diversity of medicinal and aromatic plants was studied by various workers, but the cytological study along with the meiotic abnormalities in the species has been carried out for the first time from the study area. Therefore, there is need to generate awareness among the local people regarding the conservation and utilization of medicinal plants.
View full abstractDownload PDF (724K) -
Rayan Partovi, Farah Farahani, Masoud Sheidai, Taher Nejad Satari2017 Volume 82 Issue 4 Pages 395-401
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSBananas (Musa spp.) are one of the most important fruits in the world. The mesophyll tissue of leaves excised from in vitro plantlets of banana (Musa spp.) cvs. Dwarf Cavendish and Valery were used as a source of protoplast isolation. Aliquots of purified protoplasts were plated in Murashige and Skoog (MS, 1962) liquid culture medium (F) without growth regulators. Protoplast-derived microcolonies were individually picked up and gently transferred onto regeneration medium “P” (MS medium with IAA and BAP). The highest protoplast numbers were produced in cv. Valery. Microcolonies were regenerated to plantlets, the longest mean of length shoot and the highest shoot proliferation rate were observed in cv. Dwarf Cavendish and cv. Valery, respectively. Chromosome numbers of the plants regenerated from Musa cv. Valery protoplasts varied from 66% normal (2n=3x=33), 20% (2n=2x+1=23) and 14% aneuploids (2n=2x−2=20) of the cells and 54% normal (2n=3x=33), 26% (2n=2x+1=23) and 20% aneuploids (2n=2x−2=20) of cells were in cv. Dwarf Cavendish.
View full abstractDownload PDF (572K) -
Weerayuth Supiwong, Pasakorn Saenjundaeng, Nuntiya Maneechot, Supatcha ...2017 Volume 82 Issue 4 Pages 403-411
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSA discovery of nucleolar organizer regions (NORs) polymorphism and karyological analysis in the crystal eye catfish, Hemibagrus wyckii (Bleeker, 1858) from Nong Khai and Sing Buri Provinces, Thailand, were investigated. The mitotic chromosome preparation was prepared by directly from kidney cells of five male and five female specimens. Conventional and Ag-NOR staining techniques were applied to stain the chromosomes. The results shown that the diploid chromosome number of H. wyckii was 2n=62 and the fundamental numbers (NF) of both sexes were 110. The karyotype consists of 14 large metacentric, 14 large submetacentric, 8 large acrocentric, 8 medium metacentric, 4 medium submetacentric, 6 medium telocentric, and 8 small telocentric chromosomes. No strange size chromosomes related to sex were observed. In addition, the interstitial nucleolar organizer regions (NORs) were clearly observed at the long arm of the chromosome pair No. 12. This is the first report on polymorphism of NORs in H. wyckii. The result revealed that a heteromorphic of one male and one female had a single NOR-bearing chromosome of chromosome pair No. 12 (12a12b), while four males and four females had two NOR-bearing chromosomes of the chromosome pair No. 12 with a homomorphic (12a12a). The karyotype formula for H. wyckii is as follows:
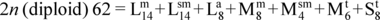 View full abstractDownload PDF (1421K)
View full abstractDownload PDF (1421K) -
Fukashi Shibata, Masahiro Hizume, Hiroaki Ohashi, Sho Furukawa2017 Volume 82 Issue 4 Pages 413-422
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSIn the complex species of Scilla scilloides involving A and B genomes, an amphidiploid having AABB genome composition is a major one of the cytotypes and distributes throughout a growing area of the species. An origin of amphidiploids and whether amphidiploids generated in monophyly or polyphyly are unsolved. In three hybrids with defied combination of genomes a maternal inheritance of chloroplast genome was confirmed in this species. An analysis of matK sequence of chloroplast genome demonstrated that a maternal parent of all amphidiploids in Japan, Korea and Taiwan investigated was a Japanese B genome diploid (BB diploid). In the analysis of ITS1and 5S rDNA of nuclear genome and genome sizes indicated that the paternal parent of amphidiploids was different between the amphidiploid from Japan and Korea, and the amphidiploid from Taiwan. The Japanese and Korean amphidiploids would be originated by a hybridization and subsequent polyploidization between the maternal parent of Japanese BB diploid and the paternal parent of Korean A genome diploid (AA diploid). Taiwanese amphidiploids were generated between the maternal parent of Japanese BB diploid and a paternal parent of AA diploid undiscovered. An event of generation of the amphidiploids in S. scilloides occurred at least twice in the different locations with the different parents.
View full abstractDownload PDF (580K) -
Kazi Nahida Begum, Sheikh Shamimul Alam2017 Volume 82 Issue 4 Pages 423-428
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSFour specimens of Gynura nepalensis DC. were investigated for authentic characterization by differential cytogenetical and PCR based molecular analysis. Three specimens out of four were collected from different localities of Bangladesh viz. i) from Netrokona district (specimen A), ii) from Dhaka district (specimen B) and iii) from Bogra district (specimen D). On the other hand, specimen C was collected from Canada. The diploid chromosome number of four specimens were 2n=20 with different centromeric formulae such as 14m+6sm in specimens A, B and D while 20m in specimen C. Total length of 2n chromosome complement was 211.61 µm in specimen A, 217.58 µm in specimen B, 168.82 µm in specimen C and 197.80 µm in specimen D. The maximum range of chromosome length was found in specimen B (6.67–14.30 µm) and minimum in specimen C (6.44–10.12 µm). The four specimens have distinct CMA- and DAPI-banding pattern which confined to the terminal regions of respective chromosomes. Karyotype analysis revealed that the number, location and distributions of GC- and AT-rich repeats are different in these four specimens. Heteromorphicity in respect of occurrence of CMA- and DAPI-banding pattern in the homologue members were observed in specimen A, C and D. The heteromorphicity might be a result of deletion of banded region from the respective homologue members. One interstitial CMA band was found on the short arm in a member of pair V in specimen A and in pair IV of specimen C. Two terminal DAPI bands were found in two chromosomes of specimens A and B. These CMA- and DAPI-banded chromosomes were unique and thus could be used as marker chromosomes for the respective specimens. Only specimen C showed RAPD bands with three primers. However, the four specimens had distinguishable ISSR-banding patterns with seven primer combinations. Specimen C showed maximum ISSR bands. The combined RAPD and ISSR analysis placed specimen C in a separate cluster with the highest genetic distance. In spite of very distant location specimens A and D were placed in a sub-cluster with narrow genetic distance.
View full abstractDownload PDF (617K) -
Chandan Kumar Dash, Mahin Afroz, Syeda Sharmeen Sultana, Sheikh Shamim ...2017 Volume 82 Issue 4 Pages 429-434
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSFour specimens of Ocimum viz. O. basilicum L., O. gratissimum L. and two forms of O. tenuiflorum L. (Green form and Purple form) were cytogenetically studied for authentic characterization. The green and purple forms of O. tenuiflorum L. are found to possess 2n=36 and 2n=34 chromosomes, respectively. On the other hand, 2n=54 chromosomes were observed in O. basilicum L. and 2n=40 chromosomes in O. gratissimum L. In this study, 2n chromosome number of O. tenuiflorum L. (Purple form) and O. basilicum L. are the first report. Six sub-metacentric chromosomes were observed in the green form of O. tenuiflorum L. (30m+6sm) while other three specimens possessed only metacentric chromosomes. These four specimens of Ocimum L. have distinct CMA- and DAPI-banding patterns based on the number, location, intensity and percentage of GC- and AT-rich repeats. Two CMA fluoresced satellites were observed in only O. gratissimum L. indicating stain specific nature of the satellite portions. Moreover, it was possible to identify certain marker chromosomes specific for each specimen with the help of CMA and DAPI banding pattern. The two forms (Green and Purple) of O. tenuiflorum L. are sharply different in respect of conventional karyotype and fluorescent banding pattern. Thus, a subtle revision is essential regarding their taxonomic status.
View full abstractDownload PDF (674K) -
Krit Pinthong, Weerayuth Supiwong, Baramate Simporn, Supatcha Choosean ...2017 Volume 82 Issue 4 Pages 435-441
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSA first report on nucleolar organizer regions (NORs) and karyological analyses in the Chevey’s sheetfish (Micronema cheveyi) from the Chao Phraya Basin, Thailand, was performed. The mitotic chromosome preparation was directly prepared from kidney cells of ten male and eight female specimens. Conventional staining and Ag-NOR banding techniques were applied to stain the chromosomes. The results shown that the diploid chromosome number of M. cheveyi was 2n=78 and the fundamental numbers (NF) were 96 in both sexes. The karyotype composed of 4 large metacentric, 6 large submetacentric, 10 large acrocentric, 32 medium telocentric, and 26 small telocentric chromosomes. No heteromorphic sex chromosomes were revealed. In addition, the telomeric NORs were clearly observed at the short arm adjacent to the telomere of the third chromosome pair (submetacentric chromosomes). The karyotype formula for M. cheveyi is as follows:
 View full abstractDownload PDF (838K)
View full abstractDownload PDF (838K) -
Mushtaq Ahmad Khah, Rakesh Chandra Verma2017 Volume 82 Issue 4 Pages 443-447
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSGamma irradiations were successfully used to develop the multiple chromosomal interchanges in pearl millet (Pennisetum glaucum L.). A plant showing two interchange complexes was isolated from the M1 population raised from 30 kR irradiated seed progeny. The translocation heterozygote showed ring and chain of one hexavalent as well as one quadrivalent along with two bivalents at diakinesis in majority of pollen mother cells. Some pollen mother cells exhibited pentavalents, trivalents and univalents in varying frequencies. The hexavalent predominantly showed alternate type of segregation when compared with the quadrivalent which exhibited adjacent type of orientation in most of the PMCs. At anaphase-I, the translocation heterozygote displayed various abnormalities such as unequal chromosome distribution, laggards and bridges. Pollen fertility was found to be very low (28.79%).
View full abstractDownload PDF (595K) -
Bundit Tengjaroenkul, Lamyai Neeratanaphan, Sitthisak Jantarat, Sitthi ...2017 Volume 82 Issue 4 Pages 449-455
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSWe report the first chromosome analysis of the Indian giant flying squirrel (Petaurista philippensis) and lesser giant flying squirrel (P. elegans) from Northeast of Thailand. Blood samples were taken from one female squirrel. After standard whole blood lymphocytes had been cultured at 37°C for 72 h in the presence of colchicine, metaphase spreads were performed on microscopic slides and air-dried. Giemsa’s staining, GTG-banding, high resolutions GTG-banding, and CBG-banding techniques were used to stain chromosomes. The results showed that the diploid chromosome number of P. philippensis and P. elegans were 2n=38 and the fundamental numbers (NF) were 76 in both species. The chromosomes presences of metacentric and submetacentric chromosomes were 12–26 and 8–28, respectively. The largest submetacentric chromosome pair 1 showed clearly observable difference size (heteromorphic, 1a1b) in both species. GTG-banding and high-resolution GTG-banding techniques showed that number of bands of P. philippensis was 127 and 166, respectively. From the CBG-banding technique of P. philippensis showed C-positive (dark band) on the centromere of all chromosomes. Moreover, we also found dark bands on telomere of the short arm of chromosome pairs 1a, 4, 6, 7 and 9. In addition, the long arm of chromosome pair 17 showed clearly observable interstitial nucleolar organizer regions (NORs). This is the first report on polymorphism of NORs in P. elegans. The result showed that a heteromorphic had a different size of NORs of chromosome pair 17 (17a17b). The karyotype formulae could be deduced as:
 View full abstractDownload PDF (1818K)
View full abstractDownload PDF (1818K) -
Kosit Sreeputhorn, Kriangsak Mangumphan, Benjawon Muanphet, Alongklod ...2017 Volume 82 Issue 4 Pages 457-463
Published: September 25, 2017
Released on J-STAGE: December 16, 2017
JOURNAL OPEN ACCESSThe first chromosomal characteristic of F1 hybrid catfish from Thailand was studied. Kidney cell samples were taken from four male and four female fish. The mitotic chromosome preparations were done directly from kidney cells. Conventional staining technique was applied to stain the chromosomes. The results showed that the diploid chromosome number of Mekong giant catfish (Pangasianodon gigas)×striped catfish (P. hypophthalmus) and spot pangasius (Pangasius larnaudii)×P. hypophthalmus were 2n=60, the fundamental number (NF) was 108 in both males and females. The chromosomes presences of metacentric, submetacentric, acrocentric, and telocentric chromosomes were 27-11-10-12 and 25-12-11-12, respectively. While the chromosomes of P. gigas, P. hypophthalmus, and P. larnaudii (parents) presences of metacentric, submetacentric, acrocentric, and telocentric chromosomes were 28-12-12-8, 26-10-8-16, and 24-14-14-8, respectively. Results obtained will increase our basic knowledge of the cytogenetics of the family Pangasiidae which could form the basis for future research and provide data to ensure their survival. The karyotype formulae could be deduced as:
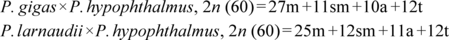 View full abstractDownload PDF (890K)
View full abstractDownload PDF (890K)
- |<
- <
- 1
- >
- >|